Evaluation of breast cancer polyclonality by combined chromosome banding and comparative genomic hybridization analysis
- PMID: 11494114
- PMCID: PMC1505591
- DOI: 10.1038/sj.neo.7900152
Evaluation of breast cancer polyclonality by combined chromosome banding and comparative genomic hybridization analysis
Abstract
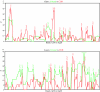
Cytogenetically unrelated clones have been detected by chromosome banding analysis in many breast carcinomas. Because these karyotypic studies were performed on short-term cultured samples, it may be argued that in vitro selection occurred or that small clones may have arisen during culturing. To address this issue, we analyzed 37 breast carcinomas by G-banding and comparative genomic hybridization (CGH), a fluorescent in situ hybridization--based screening technique that does not require culturing or tumor metaphases. All but two of the 37 karyotypically abnormal cases presented copy number changes by CGH. The picture of genomic alterations revealed by the two techniques overlapped only partly. Sometimes the CGH analysis revealed genomic imbalances that belonged to cell populations not picked up by the cytogenetic analysis and in other cases, especially when the karyotypes had many markers and chromosomes with additional material of unknown origin, CGH gave a more reliable overall picture of the copy number gains and losses. However, besides sometimes revealing cell populations with balanced chromosome aberrations or unbalanced changes that nevertheless remained undetected by CGH, G-banding analysis was essential to understand how the genomic imbalances arose in the many cases in which both techniques detected the same clonal abnormalities. Furthermore, because CGH pictures only imbalances present in a significant proportion of the test sample, the very detection by this technique of imbalances belonging to apparently small, cytogenetically unrelated clones of cells proves that these clones must have been present in vivo. This constitutes compelling evidence that the cytogenetic polyclonality observed after short-term culturing of breast carcinomas is not an artifact.
Figures
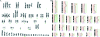

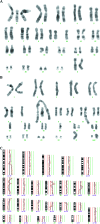

Similar articles
-
Genome profiling of breast cancer cells selected against in vitro shows copy number changes.Genes Chromosomes Cancer. 2002 Mar;33(3):304-9. doi: 10.1002/gcc.10032. Genes Chromosomes Cancer. 2002. PMID: 11807988
-
Complete cytogenetic characterization of the human breast cancer cell line MA11 combining G-banding, comparative genomic hybridization, multicolor fluorescence in situ hybridization, RxFISH, and chromosome-specific painting.Cancer Genet Cytogenet. 2001 Nov;131(1):25-30. doi: 10.1016/s0165-4608(01)00484-8. Cancer Genet Cytogenet. 2001. PMID: 11734314
-
Combined study of prostatic carcinoma by classical cytogenetic analysis and comparative genomic hybridization.Int J Oncol. 2001 Dec;19(6):1263-70. doi: 10.3892/ijo.19.6.1263. Int J Oncol. 2001. PMID: 11713598
-
Combined karyotyping, CGH and M-FISH analysis allows detailed characterization of unidentified chromosomal rearrangements in Merkel cell carcinoma.Int J Cancer. 2002 Sep 10;101(2):137-45. doi: 10.1002/ijc.10591. Int J Cancer. 2002. PMID: 12209990 Review.
-
Cytogenetic clues to breast carcinogenesis.Genes Chromosomes Cancer. 2002 Jan;33(1):1-16. doi: 10.1002/gcc.1206. Genes Chromosomes Cancer. 2002. PMID: 11746982 Review.
Cited by
-
Beyond Genetics: Metastasis as an Adaptive Response in Breast Cancer.Int J Mol Sci. 2022 Jun 3;23(11):6271. doi: 10.3390/ijms23116271. Int J Mol Sci. 2022. PMID: 35682953 Free PMC article. Review.
-
Examining metastatic behavior within 3D bioprinted vasculature for the validation of a 3D computational flow model.Sci Adv. 2020 Aug 26;6(35):eabb3308. doi: 10.1126/sciadv.abb3308. eCollection 2020 Aug. Sci Adv. 2020. PMID: 32923637 Free PMC article.
-
Use of Breast-Specific PET Scanners and Comparison with MR Imaging.Magn Reson Imaging Clin N Am. 2018 May;26(2):265-272. doi: 10.1016/j.mric.2017.12.006. Magn Reson Imaging Clin N Am. 2018. PMID: 29622131 Free PMC article. Review.
-
Profile of Chromosomal Alterations, Chromosomal Instability and Clonal Heterogeneity in Colombian Farmers Exposed to Pesticides.Front Genet. 2022 Feb 24;13:820209. doi: 10.3389/fgene.2022.820209. eCollection 2022. Front Genet. 2022. PMID: 35281828 Free PMC article.
-
Analysis of gene alterations of mitochondrial DNA D-loop regions to determine breast cancer clonality.Br J Cancer. 2012 Dec 4;107(12):2016-23. doi: 10.1038/bjc.2012.505. Epub 2012 Nov 20. Br J Cancer. 2012. PMID: 23169290 Free PMC article.
References
-
- Pandis N, Jin Y, Gorunova L, Petersson C, Bardi G, Idvall I, Johansson B, Ingvar C, Mandahl N, Mitelman F, Heim S. Chromosome analysis of 97 primary breast carcinomas: identification of eight karyotypic subgroups. Genes Chromosomes Cancer. 1995;12:173–185. - PubMed
-
- Adeyinka A, Mertens F, Idvall I, Bondeson L, Ingvar C, Mitelman F, Pandis N. Different patterns of chromosomal imbalances in metastasising and non-metastasising primary breast carcinomas. Int J Cancer. 1999;84:370–375. - PubMed
-
- Teixeira MR, Pandis N, Bardi G, Andersen JA, Mitelman F, Heim S. Clonal heterogeneity in breast cancer: karyotypic comparisons of multiple intra- and extra-tumorous samples from 3 patients. Int J Cancer. 1995;63:63–68. - PubMed
-
- Teixeira MR, Pandis N, Bardi G, Andersen JA, Heim S. Karyotypic comparisons of multiple tumorous and macroscopically normal surrounding tissue samples from patients with breast cancer. Cancer Res. 1996;56:855–859. - PubMed
-
- Kallioniemi A, Kallioniemi OP, Sudar D, Rutovitz D, Gray JW, Waldman F, Pinkel D. Comparative genomic hybridization for molecular cytogenetic analysis of solid tumors. Science. 1992;258:818–821. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
