HBV infection of cell culture: evidence for multivalent and cooperative attachment
- PMID: 11500372
- PMCID: PMC125578
- DOI: 10.1093/emboj/20.16.4443
HBV infection of cell culture: evidence for multivalent and cooperative attachment
Abstract
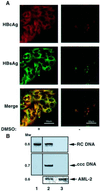
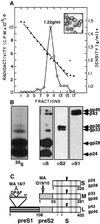
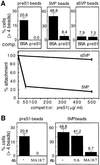
Hepadnaviruses do not infect cultured cells, therefore our knowledge of the mechanism of the early stages of virus-cell interaction is rather poor. In this study, we show that dimethylsulfoxide (DMSO)-treated HepG2 hepatoblastoma cells are infected efficiently by serum-derived hepatitis B virus (HBV) as monitored by viral gene expression and replication markers. To measure virus attachment, a variety of HBV surface proteins (HBsAgs) were conjugated to polystyrene beads and their capacity to attach cells was visualized and quantified by light microscopy at a single-cell resolution. Remarkably, DMSO increases the attachment efficiency by >200-fold. We further identify the QLDPAF sequence within preS1 as the receptor-binding viral domain epitope. Interestingly, a similar sequence is shared by several cellular, bacterial and viral proteins involved in cell adhesion, attachment and fusion. We also found that the small HBsAg contains a secondary attachment site that recognizes a distinct receptor on the cell membrane. Furthermore, we provide evidence in support of multivalent HBV attachment with synergistic interplay. Our data depict a mechanistic view of virus attachment and ingestion.
Figures






Similar articles
-
A C-terminal PreS1 sequence is sufficient to retain hepatitis B virus L protein in 293 cells.Virology. 1995 Oct 20;213(1):57-69. doi: 10.1006/viro.1995.1546. Virology. 1995. PMID: 7483279
-
Expression and purification of the synthetic preS1 gene of Hepatitis B Virus with preferred Escherichia coli codon preference.Protein Expr Purif. 2006 Jul;48(1):74-80. doi: 10.1016/j.pep.2005.11.024. Epub 2005 Dec 20. Protein Expr Purif. 2006. PMID: 16439156
-
Specific targeted binding of herpes simplex virus type 1 to hepatocytes via the human hepatitis B virus preS1 peptide.Gene Ther. 2004 Jul;11(13):1087-98. doi: 10.1038/sj.gt.3302266. Gene Ther. 2004. PMID: 15057264
-
Attachment sites and neutralising epitopes of hepatitis B virus.Minerva Gastroenterol Dietol. 2006 Mar;52(1):3-21. Minerva Gastroenterol Dietol. 2006. PMID: 16554703 Review.
-
Biological and clinical implications of HBV infection in peripheral blood mononuclear cells.Autoimmun Rev. 2008 Oct;8(1):13-7. doi: 10.1016/j.autrev.2008.07.016. Epub 2008 Aug 13. Autoimmun Rev. 2008. PMID: 18706529 Review.
Cited by
-
Quantification, epitope mapping and genotype cross-reactivity of hepatitis B preS-specific antibodies in subjects vaccinated with different dosage regimens of BM32.EBioMedicine. 2020 Sep;59:102953. doi: 10.1016/j.ebiom.2020.102953. Epub 2020 Aug 24. EBioMedicine. 2020. PMID: 32855110 Free PMC article.
-
Murine retroviral pseudotype virus containing hepatitis B virus large and small surface antigens confers specific tropism for primary human hepatocytes: a potential liver-specific targeting system.J Virol. 2002 Jan;76(2):912-7. doi: 10.1128/jvi.76.2.912-917.2002. J Virol. 2002. Retraction in: J Virol. 2019 Oct 29;93(22):e00761-19. doi: 10.1128/JVI.00761-19. PMID: 11752180 Free PMC article. Retracted.
-
Infection of a human hepatoma cell line by hepatitis B virus.Proc Natl Acad Sci U S A. 2002 Nov 26;99(24):15655-60. doi: 10.1073/pnas.232137699. Epub 2002 Nov 13. Proc Natl Acad Sci U S A. 2002. PMID: 12432097 Free PMC article.
-
Pre-structured motifs in the natively unstructured preS1 surface antigen of hepatitis B virus.Protein Sci. 2007 Oct;16(10):2108-17. doi: 10.1110/ps.072983507. Epub 2007 Aug 31. Protein Sci. 2007. PMID: 17766372 Free PMC article.
-
Replication of clinical hepatitis B virus isolate and its application for selecting antiviral agents for chronic hepatitis B patients.World J Gastroenterol. 2008 Jun 14;14(22):3490-6. doi: 10.3748/wjg.14.3490. World J Gastroenterol. 2008. PMID: 18567076 Free PMC article.
References
-
- De Meyer S., Gong,Z.J., Suwandhi,W., van Pelt,J., Soumillion,A. and Yap,S.H. (1997) Organ and species specificity of hepatitis B virus (HBV) infection: a review of literature with a special reference to preferential attachment of HBV to human hepatocytes. J. Viral Hepat., 4, 145–153. - PubMed
-
- Faktor O., De-Medina,T. and Shaul,Y. (1988) Regulation of hepatitis B virus S gene promoter in transfected cell lines. Virology, 162, 362–368. - PubMed
-
- Gripon P., Diot,C., Corlu,A. and Guguen-Guillouzo,C. (1989) Regulation by dimethylsulfoxide, insulin and corticosteroids of hepatitis B virus replication in a transfected human hepatoma cell line. J. Med. Virol., 28, 193–199. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

