Negative regulation of the Wnt-beta-catenin pathway by the transcriptional repressor HBP1
- PMID: 11500377
- PMCID: PMC125566
- DOI: 10.1093/emboj/20.16.4500
Negative regulation of the Wnt-beta-catenin pathway by the transcriptional repressor HBP1
Abstract
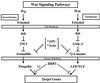
In certain cancers, constitutive Wnt signaling results from mutation in one or more pathway components. The result is the accumulation and nuclear localization of beta-catenin, which interacts with the lymphoid enhancer factor-1 (LEF)/T-cell factor (TCF) family of HMG-box transcription factors, which activate important growth regulatory genes, including cyclin D1 and c-myc. As exemplified by APC and axin, the negative regulation of beta-catenin is important for tumor suppression. Another potential mode of negative regulation is transcriptional repression of cyclin D1 and other Wnt target genes. In mammals, the transcriptional repressors in the Wnt pathway are not well defined. We have previously identified HBP1 as an HMG-box repressor and a cell cycle inhibitor. Here, we show that HBP1 is a repressor of the cyclin D1 gene and inhibits the Wnt signaling pathway. The inhibition of Wnt signaling and growth requires a common domain of HBP1. The apparent mechanism is an inhibition of TCF/LEF DNA binding through a physical interaction with HBP1. These data suggest that the suppression of Wnt signaling by HBP1 may be a mechanism to prevent inappropriate proliferation.
Figures





















Similar articles
-
Chromatin-specific regulation of LEF-1-beta-catenin transcription activation and inhibition in vitro.Genes Dev. 2001 Dec 15;15(24):3342-54. doi: 10.1101/gad.946501. Genes Dev. 2001. PMID: 11751639 Free PMC article.
-
The Yin-Yang of TCF/beta-catenin signaling.Adv Cancer Res. 2000;77:1-24. doi: 10.1016/s0065-230x(08)60783-6. Adv Cancer Res. 2000. PMID: 10549354 Review.
-
HMG box transcription factor TCF-4's interaction with CtBP1 controls the expression of the Wnt target Axin2/Conductin in human embryonic kidney cells.Nucleic Acids Res. 2003 May 1;31(9):2369-80. doi: 10.1093/nar/gkg346. Nucleic Acids Res. 2003. PMID: 12711682 Free PMC article.
-
TCF: Lady Justice casting the final verdict on the outcome of Wnt signalling.Biol Chem. 2002 Feb;383(2):255-61. doi: 10.1515/BC.2002.027. Biol Chem. 2002. PMID: 11934263 Review.
-
All Trans-Retinoic Acid Mediates MED28/HMG Box-Containing Protein 1 (HBP1)/β-Catenin Signaling in Human Colorectal Cancer Cells.J Cell Physiol. 2016 Aug;231(8):1796-803. doi: 10.1002/jcp.25285. Epub 2015 Dec 30. J Cell Physiol. 2016. PMID: 26660958
Cited by
-
HBP1-mediated Regulation of p21 Protein through the Mdm2/p53 and TCF4/EZH2 Pathways and Its Impact on Cell Senescence and Tumorigenesis.J Biol Chem. 2016 Jun 10;291(24):12688-12705. doi: 10.1074/jbc.M116.714147. Epub 2016 Apr 21. J Biol Chem. 2016. PMID: 27129219 Free PMC article.
-
MiR203 mediates subversion of stem cell properties during mammary epithelial differentiation via repression of ΔNP63α and promotes mesenchymal-to-epithelial transition.Cell Death Dis. 2013 Feb 28;4(2):e514. doi: 10.1038/cddis.2013.37. Cell Death Dis. 2013. PMID: 23449450 Free PMC article.
-
Lentivirus‑delivered nemo‑like kinase small interfering RNA inhibits laryngeal cancer cell proliferation in vitro.Mol Med Rep. 2015 Oct;12(4):5619-24. doi: 10.3892/mmr.2015.4189. Epub 2015 Aug 6. Mol Med Rep. 2015. PMID: 26252054 Free PMC article.
-
Highly specific role of the insulin receptor in breast cancer progression.Endocr Relat Cancer. 2015 Apr;22(2):145-57. doi: 10.1530/ERC-14-0490. Endocr Relat Cancer. 2015. PMID: 25694511 Free PMC article.
-
MiR-155 promotes colitis-associated intestinal fibrosis by targeting HBP1/Wnt/β-catenin signalling pathway.J Cell Mol Med. 2021 May;25(10):4765-4775. doi: 10.1111/jcmm.16445. Epub 2021 Mar 26. J Cell Mol Med. 2021. PMID: 33769664 Free PMC article.
References
-
- Albanese C.J., Johnson,G., Watanabe,N., Eklund,D., Vu,D. and Pestell,R.G. (1995) Transforming p21 ras mutants and ets2 activate the cyclin D1 promoter through distinguishable regions. J. Biol. Chem., 270, 23589–23597. - PubMed
-
- Barker N., Morin,P.J. and Clevers,H. (2000) The yin–yang of TCF/β-catenin signaling. Adv. Cancer Res., 77, 1–24. - PubMed
-
- Bieche I., Khodja,A., Driouch,K. and Lidereau,R. (1997) Genetic alteration mapping on chromosome 7 in primary breast cancer. Clin. Cancer Res., 3, 1009–1016. - PubMed
-
- Bienz M. (1998) TCF: transcriptional activator or repressor. Curr. Opin. Cell Biol., 10, 366–372. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

