Independent and coordinate effects of ADP-ribosyltransferase and GTPase-activating activities of exoenzyme S on HT-29 epithelial cell function
- PMID: 11500401
- PMCID: PMC98641
- DOI: 10.1128/IAI.69.9.5318-5328.2001
Independent and coordinate effects of ADP-ribosyltransferase and GTPase-activating activities of exoenzyme S on HT-29 epithelial cell function
Abstract
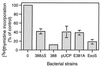
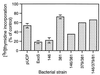
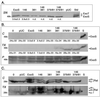
Type III-mediated translocation of exoenzyme S (ExoS) into HT-29 epithelial cells by Pseudomonas aeruginosa causes complex alterations in cell function, including inhibition of DNA synthesis, altered cytoskeletal structure, loss of readherence, microvillus effacement, and interruption of signal transduction. ExoS is a bifunctional protein having both GTPase-activating (GAP) and ADP-ribosyltransferase (ADPRT) functional domains. Comparisons of alterations in HT-29 cell function caused by P. aeruginosa strains that translocate ExoS having GAP or ADPRT mutations allowed the independent and coordinate functions of the two activities to be assessed. An E381A ADPRT mutation revealed that ExoS ADPRT activity was required for effects of ExoS on DNA synthesis and long-term cell rounding. Conversely, the R146A GAP mutation appeared to have little impact on the cellular effects of ExoS. While transient cell rounding was detected following exposure to the E381A mutant, this rounding was eliminated by an E379A-E381A ADPRT double mutation, implying that residual ADPRT activity, rather than GAP activity, was effecting transient cell rounding by the E381A mutant. To explore this possibility, E381A and R146A-E381A mutants were examined for their ability to ADP-ribosylate Ras in vitro or in vivo. While no ADP-ribosylation of Ras was detected by either mutant in vitro, both mutants were able to modify Ras when translocated by the bacteria, with the R146A-E381A mutant causing more efficient modification than the E381A mutant, in association with increased inhibition of DNA synthesis. Comparisons of Ras ADP-ribosylation by wild-type and E381A mutant ExoS by two-dimensional electrophoresis found the former to ADP-ribosylate Ras at two sites, while the latter modified Ras only once. These studies draw attention to the key role of ExoS ADPRT activity in causing the effects of bacterially translocated ExoS on DNA synthesis and cell rounding. In addition, the studies provide insight into the enhancement of ExoS ADPRT activity within the eukaryotic cell microenvironment and into possible modulatory roles that the GAP and ADPRT domains might have on the function of each other.
Figures
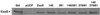




Similar articles
-
Examination of the coordinate effects of Pseudomonas aeruginosa ExoS on Rac1.Infect Immun. 2005 Sep;73(9):5458-67. doi: 10.1128/IAI.73.9.5458-5467.2005. Infect Immun. 2005. PMID: 16113262 Free PMC article.
-
ADP-ribosylation and functional effects of Pseudomonas exoenzyme S on cellular RalA.Biochemistry. 2002 Jul 30;41(30):9680-7. doi: 10.1021/bi025826n. Biochemistry. 2002. PMID: 12135390
-
Modification of Ras in eukaryotic cells by Pseudomonas aeruginosa exoenzyme S.Infect Immun. 1998 Jun;66(6):2607-13. doi: 10.1128/IAI.66.6.2607-2613.1998. Infect Immun. 1998. PMID: 9596723 Free PMC article.
-
Pseudomonas aeruginosa exoenzyme S, a bifunctional type-III secreted cytotoxin.Int J Med Microbiol. 2000 Oct;290(4-5):381-7. doi: 10.1016/S1438-4221(00)80047-8. Int J Med Microbiol. 2000. PMID: 11111915 Review.
-
Pseudomonas aeruginosa ExoS and ExoT.Rev Physiol Biochem Pharmacol. 2004;152:79-92. doi: 10.1007/s10254-004-0031-7. Epub 2004 Aug 24. Rev Physiol Biochem Pharmacol. 2004. PMID: 15375697 Review.
Cited by
-
Acquisition and evolution of the exoU locus in Pseudomonas aeruginosa.J Bacteriol. 2006 Jun;188(11):4037-50. doi: 10.1128/JB.02000-05. J Bacteriol. 2006. PMID: 16707695 Free PMC article.
-
Pseudomonas aeruginosa infection of airway epithelial cells modulates expression of Kruppel-like factors 2 and 6 via RsmA-mediated regulation of type III exoenzymes S and Y.Infect Immun. 2006 Oct;74(10):5893-902. doi: 10.1128/IAI.00489-06. Infect Immun. 2006. PMID: 16988269 Free PMC article.
-
The Role of Pseudomonas aeruginosa Virulence Factors in Cytoskeletal Dysregulation and Lung Barrier Dysfunction.Toxins (Basel). 2021 Nov 2;13(11):776. doi: 10.3390/toxins13110776. Toxins (Basel). 2021. PMID: 34822560 Free PMC article. Review.
-
Pseudomonas aeruginosa Takes a Multi-Target Approach to Achieve Junction Breach.Front Cell Infect Microbiol. 2018 Jan 11;7:532. doi: 10.3389/fcimb.2017.00532. eCollection 2017. Front Cell Infect Microbiol. 2018. PMID: 29379773 Free PMC article. Review.
-
Role of host cell polarity and leading edge properties in Pseudomonas type III secretion.Microbiology (Reading). 2010 Feb;156(Pt 2):356-373. doi: 10.1099/mic.0.033241-0. Epub 2009 Nov 12. Microbiology (Reading). 2010. PMID: 19910414 Free PMC article.
References
-
- Coburn J, Wyatt R T, Iglewski B H, Gill D M. Several GTP-binding proteins, including p21c-H-ras, are preferred substrates of Pseudomonas aeruginosa exoenzyme S. J Biol Chem. 1989;264:9004–9008. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

