Chlamydia pneumoniae expresses genes required for DNA replication but not cytokinesis during persistent infection of HEp-2 cells
- PMID: 11500413
- PMCID: PMC98653
- DOI: 10.1128/IAI.69.9.5423-5429.2001
Chlamydia pneumoniae expresses genes required for DNA replication but not cytokinesis during persistent infection of HEp-2 cells
Abstract
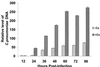
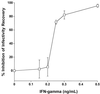
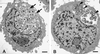
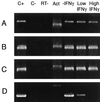
Chlamydia pneumoniae causes community-acquired pneumonia and is associated with several chronic diseases, including asthma and atherosclerosis. The intracellular growth rate of C. pneumoniae slows dramatically during chronic infection, and such persistence leads to attenuated production of new elementary bodies, appearance of morphologically aberrant reticulate bodies, and altered expression of several chlamydial genes. We used an in vitro system to further characterize persistent C. pneumoniae infection, employing both ultrastructural and transcriptional activity measurements. HEp-2 cells were infected with C. pneumoniae (TW-183) at a multiplicity of infection of 3:1, and at 2 h postinfection gamma interferon (IFN-gamma) was added to the medium at 0.15 or 0.50 ng/ml. Treated and untreated cultures were harvested at several times postinfection. RNA was isolated and reverse transcribed, and reverse transcription (RT)-PCR analyses targeting primary transcripts from chlamydial rRNA operons as well as dnaA, polA, mutS, minD, ftsK, and ftsW mRNA were done. Some cultures were fixed and stained for electron microscopic analysis, and a real-time PCR assay was used to assess relative chlamydial chromosome accumulation under each culture condition. The latter assays showed that bacterial chromosome copies accumulated severalfold during IFN-gamma treatment of infected HEp-2 cells, although less accumulation was observed in cells treated with the higher dose. Electron microscopy demonstrated that high-dose IFN-gamma treatment elicited aberrant forms of the bacterium. RT-PCR showed that chlamydial primary rRNA transcripts were present in all IFN-gamma-treated and untreated cell cultures, indicating bacterial metabolic activity. Transcripts from dnaA, polA, mutS, and minD, all of which encode products for bacterial chromosome replication and partition, were expressed in IFN-gamma-treated and untreated cells. In contrast, ftsK and ftsW, encoding products for bacterial cell division, were expressed in untreated cells, but expression was attenuated in cells treated with low-dose IFN-gamma and absent in cells given the high dose of cytokine. Thus, the development of persistence included production of transcripts for DNA replication-related, but not cell division-related, genes. These results provide new insight regarding molecular activities that accompany persistence of C. pneumoniae, as well as suggesting requirements for reactivation from persistent to productive growth.
Figures


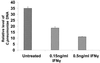
Similar articles
-
Expression of Chlamydia trachomatis genes encoding products required for DNA synthesis and cell division during active versus persistent infection.Mol Microbiol. 2001 Aug;41(3):731-41. doi: 10.1046/j.1365-2958.2001.02550.x. Mol Microbiol. 2001. PMID: 11532140
-
IFN-gamma induced persistent Chlamydia pneumoniae infection in HL and Mono Mac 6 cells: characterization by real-time quantitative PCR and culture.Microb Pathog. 2004 Jan;36(1):41-50. doi: 10.1016/j.micpath.2003.09.001. Microb Pathog. 2004. PMID: 14643639
-
Chlamydophila (Chlamydia) pneumoniae infection of human astrocytes and microglia in culture displays an active, rather than a persistent, phenotype.Am J Med Sci. 2006 Oct;332(4):168-74. doi: 10.1097/00000441-200610000-00003. Am J Med Sci. 2006. PMID: 17031241
-
The intracellular life of chlamydiae.Semin Pediatr Infect Dis. 2002 Oct;13(4):239-48. doi: 10.1053/spid.2002.127201. Semin Pediatr Infect Dis. 2002. PMID: 12491229 Review.
-
[Chlamydia pneumoniae infections].Kekkaku. 2006 Sep;81(9):581-8. Kekkaku. 2006. PMID: 17037392 Review. Japanese.
Cited by
-
Mixed infections with Chlamydia and porcine epidemic diarrhea virus - a new in vitro model of chlamydial persistence.BMC Microbiol. 2010 Jul 27;10:201. doi: 10.1186/1471-2180-10-201. BMC Microbiol. 2010. PMID: 20663197 Free PMC article.
-
Tryptophan Codon-Dependent Transcription in Chlamydia pneumoniae during Gamma Interferon-Mediated Tryptophan Limitation.Infect Immun. 2016 Aug 19;84(9):2703-13. doi: 10.1128/IAI.00377-16. Print 2016 Sep. Infect Immun. 2016. PMID: 27400720 Free PMC article.
-
Gene expression profiles of Chlamydophila pneumoniae during the developmental cycle and iron depletion-mediated persistence.PLoS Pathog. 2007 Jun;3(6):e83. doi: 10.1371/journal.ppat.0030083. PLoS Pathog. 2007. PMID: 17590080 Free PMC article.
-
Protein expression profiles of Chlamydia pneumoniae in models of persistence versus those of heat shock stress response.Infect Immun. 2006 Jul;74(7):3853-63. doi: 10.1128/IAI.02104-05. Infect Immun. 2006. PMID: 16790757 Free PMC article.
-
Chlamydia Persistence: A Survival Strategy to Evade Antimicrobial Effects in-vitro and in-vivo.Front Microbiol. 2018 Dec 12;9:3101. doi: 10.3389/fmicb.2018.03101. eCollection 2018. Front Microbiol. 2018. PMID: 30619180 Free PMC article. Review.
References
-
- Blythe M J, Katz B P, Batteiger B E, Ganser J A, Jones R B. Recurrent genitourinary chlamydial infection in sexually active female adolescents. J Pediatr. 1992;121:487–493. - PubMed
-
- Donachie W D. The cell cycle of Escherichia coli. Nucleic Acids Res. 1993;18:6713–6721.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

