Novel anther-specific myb genes from tobacco as putative regulators of phenylalanine ammonia-lyase expression
- PMID: 11500571
- PMCID: PMC117172
- DOI: 10.1104/pp.126.4.1738
Novel anther-specific myb genes from tobacco as putative regulators of phenylalanine ammonia-lyase expression
Abstract
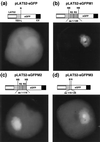
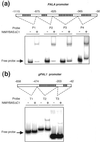
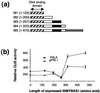
Two cDNA clones (NtmybAS1 and NtmybAS2) encoding MYB-related proteins with strong sequence similarity to petunia (Petunia hybrida) PhMYB3 were isolated from a tobacco (Nicotiana tabacum cv Samsun) pollen cDNA library. Northern blot and in situ hybridization revealed that NtmybAS transcripts are specifically expressed in both sporophytic and gametophytic tissues of the anther including tapetum, stomium, vascular tissue, and developing pollen. Random binding site selection assays revealed that NtMYBAS1 bound to DNA sequences closely resembling consensus MYB binding sites MBSI and MBSIIG, with a higher affinity for MBSI. Transient expression analyses of the N-terminal MYB domain demonstrated the presence of functional nuclear localization signals, and full-length NtMYBAS1 was able to activate two different phenylalanine ammonia-lyase promoters (PALA and gPAL1) in tobacco leaf protoplasts. Similar analysis of truncated NtmybAS1 cDNAs identified an essential, C-terminal trans-activation domain. Further in situ hybridization analyses demonstrated strict co-expression of NtmybAS and gPAL1 in the tapetum and stomium. Despite abundant expression of NtmybAS transcripts in mature pollen, gPAL1 transcripts were not detectable in pollen. Our data demonstrate that NtMYBAS1 is a functional anther-specific transcription factor, which is likely to be a positive regulator of gPAL1 expression and phenylpropanoid synthesis in sporophytic, but not in gametophytic, tissues of the anther.
Figures








Similar articles
-
Phenylalanine ammonia-lyase gene structure, expression, and evolution in Nicotiana.Plant Mol Biol. 1996 Feb;30(4):711-22. doi: 10.1007/BF00019006. Plant Mol Biol. 1996. PMID: 8624404
-
Phenylalanine ammonia-lyase in tobacco. Molecular cloning and gene expression during the hypersensitive reaction to tobacco mosaic virus and the response to a fungal elicitor.Plant Physiol. 1994 Nov;106(3):877-86. doi: 10.1104/pp.106.3.877. Plant Physiol. 1994. PMID: 7824656 Free PMC article.
-
The tobacco bZIP transcription factor BZI-1 binds to G-box elements in the promoters of phenylpropanoid pathway genes in vitro, but it is not involved in their regulation in vivo.Mol Genet Genomics. 2002 Mar;267(1):16-26. doi: 10.1007/s00438-001-0636-3. Epub 2002 Feb 12. Mol Genet Genomics. 2002. PMID: 11919711
-
Self-incompatibility: a self-recognition system in plants.Science. 1990 Nov 16;250(4983):937-41. doi: 10.1126/science.2237440. Science. 1990. PMID: 2237440 Review.
-
Methylidene-imidazolone (MIO) from histidine and phenylalanine ammonia-lyase.Adv Protein Chem. 2001;58:175-214. doi: 10.1016/s0065-3233(01)58005-5. Adv Protein Chem. 2001. PMID: 11665488 Review. No abstract available.
Cited by
-
EMB2473/MIRO1, an Arabidopsis Miro GTPase, is required for embryogenesis and influences mitochondrial morphology in pollen.Plant Cell. 2008 Mar;20(3):589-601. doi: 10.1105/tpc.107.055756. Epub 2008 Mar 14. Plant Cell. 2008. PMID: 18344283 Free PMC article.
-
Investigating Triticeae anther gene promoter activity in transgenic Brachypodium distachyon.Planta. 2017 Feb;245(2):385-396. doi: 10.1007/s00425-016-2612-5. Epub 2016 Oct 27. Planta. 2017. PMID: 27787603
-
The MADS box transcription factor ZmMADS2 is required for anther and pollen maturation in maize and accumulates in apoptotic bodies during anther dehiscence.Plant Physiol. 2004 Mar;134(3):1069-79. doi: 10.1104/pp.103.030577. Epub 2004 Mar 4. Plant Physiol. 2004. PMID: 15001699 Free PMC article.
-
AthaMap: an online resource for in silico transcription factor binding sites in the Arabidopsis thaliana genome.Nucleic Acids Res. 2004 Jan 1;32(Database issue):D368-72. doi: 10.1093/nar/gkh017. Nucleic Acids Res. 2004. PMID: 14681436 Free PMC article.
-
MYB transcription factors-master regulators of phenylpropanoid biosynthesis and diverse developmental and stress responses.Plant Cell Rep. 2022 Dec;41(12):2245-2260. doi: 10.1007/s00299-022-02927-1. Epub 2022 Sep 29. Plant Cell Rep. 2022. PMID: 36171500 Review.
References
-
- Aarts MG, Hodge R, Kalantidis K, Florack D, Wilson ZA, Mulligan BJ, Stiekema WJ, Scott R, Pereira A. The Arabidopsis MALE STERILITY 2 protein shares similarity with reductases in elongation/condensation complexes. Plant J. 1997;12:615–623. - PubMed
-
- Avila J, Nieto C, Canas L, Benito MJ, Paz-Ares J. Petunia hybrida genes related to the maize regulatory C1 gene and to animal myb proto-oncogenes. Plant J. 1993;3:553–562. - PubMed
-
- Baltz R, Domon C, Pillay DT, Steinmetz A. Characterization of a pollen-specific cDNA from sunflower encoding a zinc finger protein. Plant J. 1992;2:713–721. - PubMed
-
- Bate NJ, Orr J, Ni W, Meromi A, Nadler-Hassar T, Doerner PW, Dixon RA, Lamb CJ, Elkind Y. Quantitative relationship between phenylalanine ammonia-lyase levels and phenylpropanoid accumulation in transgenic tobacco identifies a rate-determining step in natural product synthesis. Proc Natl Acad Sci USA. 1994;91:7608–7612. - PMC - PubMed
-
- Blackwell TK, Kretzner L, Blackwood EM, Eisenman RN, Weintraub H. Sequence-specific DNA binding by the c-Myc protein. Science. 1990;23:1149–1151. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

