Structural and functional interactions of transcription factor (TF) IIA with TFIIE and TFIIF in transcription initiation by RNA polymerase II
- PMID: 11509574
- PMCID: PMC4492724
- DOI: 10.1074/jbc.M106422200
Structural and functional interactions of transcription factor (TF) IIA with TFIIE and TFIIF in transcription initiation by RNA polymerase II
Abstract
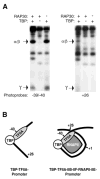
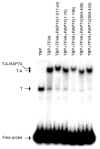
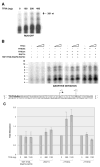
A topological model for transcription initiation by RNA polymerase II (RNAPII) has recently been proposed. This model stipulates that wrapping of the promoter DNA around RNAPII and the general initiation factors TBP, TFIIB, TFIIE, TFIIF and TFIIH induces a torsional strain in the DNA double helix that facilitates strand separation and open complex formation. In this report, we show that TFIIA, a factor previously shown to both stimulate basal transcription and have co-activator functions, is located near the cross-point of the DNA loop where it can interact with TBP, TFIIE56, TFIIE34, and the RNAPII-associated protein (RAP) 74. In addition, we demonstrate that TFIIA can stimulate basal transcription by stimulating the functions of both TFIIE34 and RAP74 during the initiation step of the transcription reaction. These results provide novel insights into mechanisms of TFIIA function.
Figures



Similar articles
-
Localization of subunits of transcription factors IIE and IIF immediately upstream of the transcriptional initiation site of the adenovirus major late promoter.J Biol Chem. 1996 Apr 12;271(15):8517-20. doi: 10.1074/jbc.271.15.8517. J Biol Chem. 1996. PMID: 8621472
-
Topological localization of the human transcription factors IIA, IIB, TATA box-binding protein, and RNA polymerase II-associated protein 30 on a class II promoter.J Biol Chem. 1994 Aug 5;269(31):19962-7. J Biol Chem. 1994. PMID: 8051080
-
Wrapping of promoter DNA around the RNA polymerase II initiation complex induced by TFIIF.Mol Cell. 1998 Sep;2(3):341-51. doi: 10.1016/s1097-2765(00)80278-6. Mol Cell. 1998. PMID: 9774972 Free PMC article.
-
Conserved RNA polymerase II initiation complex structure.Curr Opin Struct Biol. 2017 Dec;47:17-22. doi: 10.1016/j.sbi.2017.03.013. Epub 2017 Apr 22. Curr Opin Struct Biol. 2017. PMID: 28437704 Review.
-
Structural Insights into the Eukaryotic Transcription Initiation Machinery.Annu Rev Biophys. 2017 May 22;46:59-83. doi: 10.1146/annurev-biophys-070816-033751. Annu Rev Biophys. 2017. PMID: 28532216 Free PMC article. Review.
Cited by
-
Characterization of a multisubunit transcription factor complex essential for spliced-leader RNA gene transcription in Trypanosoma brucei.Mol Cell Biol. 2005 Aug;25(16):7303-13. doi: 10.1128/MCB.25.16.7303-7313.2005. Mol Cell Biol. 2005. PMID: 16055738 Free PMC article.
-
Full and partial genome-wide assembly and disassembly of the yeast transcription machinery in response to heat shock.Genes Dev. 2006 Aug 15;20(16):2250-65. doi: 10.1101/gad.1437506. Genes Dev. 2006. PMID: 16912275 Free PMC article.
-
Photo-cross-linking of a purified preinitiation complex reveals central roles for the RNA polymerase II mobile clamp and TFIIE in initiation mechanisms.Mol Cell Biol. 2004 Feb;24(3):1122-31. doi: 10.1128/MCB.24.3.1122-1131.2004. Mol Cell Biol. 2004. PMID: 14729958 Free PMC article.
-
Functional dissection of the catalytic mechanism of mammalian RNA polymerase II.Biochem Cell Biol. 2005 Aug;83(4):497-504. doi: 10.1139/o05-061. Biochem Cell Biol. 2005. PMID: 16094453 Free PMC article. Review.
-
The highly conserved glutamic acid 791 of Rpb2 is involved in the binding of NTP and Mg(B) in the active center of human RNA polymerase II.Nucleic Acids Res. 2005 May 10;33(8):2629-39. doi: 10.1093/nar/gki570. Print 2005. Nucleic Acids Res. 2005. PMID: 15886393 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

