The loop domain of heat shock transcription factor 1 dictates DNA-binding specificity and responses to heat stress
- PMID: 11511544
- PMCID: PMC312766
- DOI: 10.1101/gad.894801
The loop domain of heat shock transcription factor 1 dictates DNA-binding specificity and responses to heat stress
Abstract
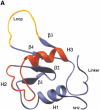
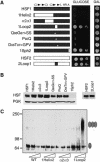
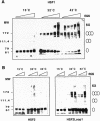
Eukaryotic heat shock transcription factors (HSF) regulate an evolutionarily conserved stress-response pathway essential for survival against a variety of environmental and developmental stresses. Although the highly similar HSF family members have distinct roles in responding to stress and activating target gene expression, the mechanisms that govern these roles are unknown. Here we identify a loop within the HSF1 DNA-binding domain that dictates HSF isoform specific DNA binding in vitro and preferential target gene activation by HSF family members in both a yeast transcription assay and in mammalian cells. These characteristics of the HSF1 loop region are transposable to HSF2 and sufficient to confer DNA-binding specificity, heat shock inducible HSP gene expression and protection from heat-induced apoptosis in vivo. In addition, the loop suppresses formation of the HSF1 trimer under basal conditions and is required for heat-inducible trimerization in a purified system in vitro, suggesting that this domain is a critical part of the HSF1 heat-stress-sensing mechanism. We propose that this domain defines a signature for HSF1 that constitutes an important determinant for how cells utilize a family of transcription factors to respond to distinct stresses.
Figures







References
-
- Abravaya K, Myers MP, Murphy SP, Morimoto RI. The human heat shock protein hsp70 interacts with HSF, the transcription factor that regulates heat shock gene expression. Genes & Dev. 1992;6:1153–1164. - PubMed
-
- Christians E, Davis AA, Thomas SD, Benjamin IJ. Maternal effect of Hsf1 on reproductive success. Nature. 2000;407:693–694. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
