Alginate overproduction affects Pseudomonas aeruginosa biofilm structure and function
- PMID: 11514525
- PMCID: PMC95424
- DOI: 10.1128/JB.183.18.5395-5401.2001
Alginate overproduction affects Pseudomonas aeruginosa biofilm structure and function
Abstract
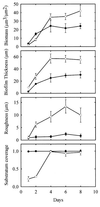
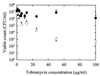
During the course of chronic cystic fibrosis (CF) infections, Pseudomonas aeruginosa undergoes a conversion to a mucoid phenotype, which is characterized by overproduction of the exopolysaccharide alginate. Chronic P. aeruginosa infections involve surface-attached, highly antibiotic-resistant communities of microorganisms organized in biofilms. Although biofilm formation and the conversion to mucoidy are both important aspects of CF pathogenesis, the relationship between them is at the present unclear. In this study, we report that the overproduction of alginate affects biofilm development on an abiotic surface. Biofilms formed by an alginate-overproducing strain exhibit a highly structured architecture and are significantly more resistant to the antibiotic tobramycin than a biofilm formed by an isogenic nonmucoid strain. These results suggest that an important consequence of the conversion to mucoidy is an altered biofilm architecture that shows increasing resistance to antimicrobial treatments.
Figures



References
-
- Christensen B B, Sternberg C, Andersen J B, Palmer R J, Jr, Nielsen A T, Givskov M, Molin S. Molecular tools for study of biofilm physiology. Methods Enzymol. 1999;310:20–42. - PubMed
-
- Costerton J W, Lewandowski Z, Caldwell D E, Korber D R, Lappin-Scott H M. Microbial biofilms. Annu Rev Microbiol. 1995;49:711–745. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

