ERB1, the yeast homolog of mammalian Bop1, is an essential gene required for maturation of the 25S and 5.8S ribosomal RNAs
- PMID: 11522832
- PMCID: PMC55883
- DOI: 10.1093/nar/29.17.3621
ERB1, the yeast homolog of mammalian Bop1, is an essential gene required for maturation of the 25S and 5.8S ribosomal RNAs
Abstract
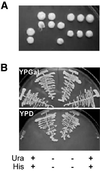
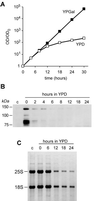
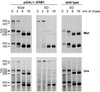
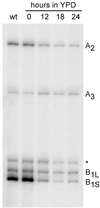
We have recently shown that the mammalian nucleolar protein Bop1 is involved in synthesis of the 28S and 5.8S ribosomal RNAs (rRNAs) and large ribosome subunits in mouse cells. Here we have investigated the functions of the Saccharomyces cerevisiae homolog of Bop1, Erb1p, encoded by the previously uncharacterized open reading frame YMR049C. Gene disruption showed that ERB1 is essential for viability. Depletion of Erb1p resulted in a loss of 25S and 5.8S rRNAs synthesis, while causing only a moderate reduction and not a complete block in 18S rRNA formation. Processing analysis showed that Erb1p is required for synthesis of 7S pre-rRNA and mature 25S rRNA from 27SB pre-rRNA. In Erb1p-depleted cells these products of 27SB processing are largely absent and 27SB pre-rRNA is under-accumulated, apparently due to degradation. In addition, depletion of Erb1p caused delayed processing of the 35S pre-rRNA. These findings demonstrate that Erb1p, like its mammalian counterpart Bop1, is required for formation of rRNA components of the large ribosome particles. The similarities in processing defects caused by functional disruption of Erb1p and Bop1 suggest that late steps in maturation of the large ribosome subunit rRNAs employ mechanisms that are evolutionarily conserved throughout eukaryotes.
Figures



Similar articles
-
Bop1 is a mouse WD40 repeat nucleolar protein involved in 28S and 5. 8S RRNA processing and 60S ribosome biogenesis.Mol Cell Biol. 2000 Aug;20(15):5516-28. doi: 10.1128/MCB.20.15.5516-5528.2000. Mol Cell Biol. 2000. PMID: 10891491 Free PMC article.
-
The exosome subunit Rrp43p is required for the efficient maturation of 5.8S, 18S and 25S rRNA.Nucleic Acids Res. 1999 Mar 1;27(5):1283-8. doi: 10.1093/nar/27.5.1283. Nucleic Acids Res. 1999. PMID: 9973615 Free PMC article.
-
The 3' end of yeast 5.8S rRNA is generated by an exonuclease processing mechanism.Genes Dev. 1996 Feb 15;10(4):502-13. doi: 10.1101/gad.10.4.502. Genes Dev. 1996. PMID: 8600032
-
Mak16p is required for the maturation of 25S and 5.8S rRNAs in the yeast Saccharomyces cerevisiae.Yeast. 2006 May;23(7):495-506. doi: 10.1002/yea.1368. Yeast. 2006. PMID: 16710831
-
IT'S 2 for the price of 1: Multifaceted ITS2 processing machines in RNA and DNA maintenance.DNA Repair (Amst). 2019 Sep;81:102653. doi: 10.1016/j.dnarep.2019.102653. Epub 2019 Jul 8. DNA Repair (Amst). 2019. PMID: 31324529 Free PMC article. Review.
Cited by
-
Rrp5p, Noc1p and Noc2p form a protein module which is part of early large ribosomal subunit precursors in S. cerevisiae.Nucleic Acids Res. 2013 Jan;41(2):1191-210. doi: 10.1093/nar/gks1056. Epub 2012 Dec 2. Nucleic Acids Res. 2013. PMID: 23209026 Free PMC article.
-
Assembly of Saccharomyces cerevisiae 60S ribosomal subunits: role of factors required for 27S pre-rRNA processing.EMBO J. 2011 Sep 16;30(19):4020-32. doi: 10.1038/emboj.2011.338. EMBO J. 2011. PMID: 21926967 Free PMC article.
-
The carboxy-terminal domain of Erb1 is a seven-bladed ß-propeller that binds RNA.PLoS One. 2015 Apr 16;10(4):e0123463. doi: 10.1371/journal.pone.0123463. eCollection 2015. PLoS One. 2015. PMID: 25880847 Free PMC article.
-
Quality control mechanisms during ribosome maturation.Trends Cell Biol. 2013 May;23(5):242-50. doi: 10.1016/j.tcb.2013.01.004. Epub 2013 Feb 1. Trends Cell Biol. 2013. PMID: 23375955 Free PMC article. Review.
-
Ytm1, Nop7, and Erb1 form a complex necessary for maturation of yeast 66S preribosomes.Mol Cell Biol. 2005 Dec;25(23):10419-32. doi: 10.1128/MCB.25.23.10419-10432.2005. Mol Cell Biol. 2005. PMID: 16287855 Free PMC article.
References
-
- Venema J. and Tollervey,D. (1999) Ribosome synthesis in Saccharomyces cerevisiae. Annu. Rev. Genet., 33, 261–311. - PubMed
-
- Eichler D.C. and Craig,N. (1994) Processing of eukaryotic ribosomal RNA. Prog. Nucleic Acid Res. Mol. Biol., 49, 197–239. - PubMed
-
- Neer E.J., Schmidt,C.J., Nambudripad,R. and Smith,T.F. (1994) The ancient regulatory-protein family of WD-repeat proteins. Nature, 371, 297–300. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

