Structure of the ABC ATPase domain of human TAP1, the transporter associated with antigen processing
- PMID: 11532960
- PMCID: PMC125601
- DOI: 10.1093/emboj/20.17.4964
Structure of the ABC ATPase domain of human TAP1, the transporter associated with antigen processing
Abstract
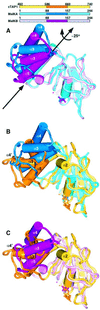
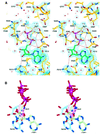
The transporter associated with antigen processing (TAP) is an ABC transporter formed of two subunits, TAP1 and TAP2, each of which has an N-terminal membrane-spanning domain and a C-terminal ABC ATPase domain. We report the structure of the C-terminal ABC ATPase domain of TAP1 (cTAP1) bound to ADP. cTAP1 forms an L-shaped molecule with two domains, a RecA-like domain and a small alpha-helical domain. The diphosphate group of ADP interacts with the P-loop as expected. Residues thought to be involved in gamma-phosphate binding and hydrolysis show flexibility in the ADP-bound state as evidenced by their high B-factors. Comparisons of cTAP1 with other ABC ATPases from the ABC transporter family as well as ABC ATPases involved in DNA maintenance and repair reveal key regions and residues specific to each family. Three ATPase subfamilies are identified which have distinct adenosine recognition motifs, as well as distinct subdomains that may be specific to the different functions of each subfamily. Differences between TAP1 and TAP2 in the nucleotide-binding site may be related to the observed asymmetry during peptide transport.
Figures



Similar articles
-
The distinct nucleotide binding states of the transporter associated with antigen processing (TAP) are regulated by the nonhomologous C-terminal tails of TAP1 and TAP2.Eur J Biochem. 2003 Nov;270(22):4531-46. doi: 10.1046/j.1432-1033.2003.03848.x. Eur J Biochem. 2003. PMID: 14622282
-
Analyses of conformational states of the transporter associated with antigen processing (TAP) protein in a native cellular membrane environment.J Biol Chem. 2013 Dec 27;288(52):37039-47. doi: 10.1074/jbc.M113.504696. Epub 2013 Nov 6. J Biol Chem. 2013. PMID: 24196954 Free PMC article.
-
A half-type ABC transporter TAPL is highly conserved between rodent and man, and the human gene is not responsive to interferon-gamma in contrast to TAP1 and TAP2.J Biochem. 2000 Oct;128(4):711-8. doi: 10.1093/oxfordjournals.jbchem.a022805. J Biochem. 2000. PMID: 11011155
-
ABC-ATPases, adaptable energy generators fuelling transmembrane movement of a variety of molecules in organisms from bacteria to humans.J Mol Biol. 1999 Oct 22;293(2):381-99. doi: 10.1006/jmbi.1999.2993. J Mol Biol. 1999. PMID: 10529352 Review.
-
How does TAP pump peptides? insights from DNA repair and traffic ATPases.Immunol Today. 2000 Dec;21(12):598-600. doi: 10.1016/s0167-5699(00)01720-5. Immunol Today. 2000. PMID: 11114418 Review. No abstract available.
Cited by
-
Characterization of the amicetin biosynthesis gene cluster from Streptomyces vinaceusdrappus NRRL 2363 implicates two alternative strategies for amide bond formation.Appl Environ Microbiol. 2012 Apr;78(7):2393-401. doi: 10.1128/AEM.07185-11. Epub 2012 Jan 20. Appl Environ Microbiol. 2012. PMID: 22267658 Free PMC article.
-
Direct evidence that the N-terminal extensions of the TAP complex act as autonomous interaction scaffolds for the assembly of the MHC I peptide-loading complex.Cell Mol Life Sci. 2012 Oct;69(19):3317-27. doi: 10.1007/s00018-012-1005-6. Epub 2012 May 27. Cell Mol Life Sci. 2012. PMID: 22638925 Free PMC article.
-
Structure of adeno-associated virus type 2 Rep40-ADP complex: insight into nucleotide recognition and catalysis by superfamily 3 helicases.Proc Natl Acad Sci U S A. 2004 Aug 24;101(34):12455-60. doi: 10.1073/pnas.0403454101. Epub 2004 Aug 13. Proc Natl Acad Sci U S A. 2004. PMID: 15310852 Free PMC article.
-
Mutations in the linker domain of NBD2 of SUR inhibit transduction but not nucleotide binding.EMBO J. 2002 Aug 15;21(16):4250-8. doi: 10.1093/emboj/cdf419. EMBO J. 2002. PMID: 12169627 Free PMC article.
-
Side chain and backbone contributions of Phe508 to CFTR folding.Nat Struct Mol Biol. 2005 Jan;12(1):10-6. doi: 10.1038/nsmb881. Epub 2004 Dec 26. Nat Struct Mol Biol. 2005. PMID: 15619636 Free PMC article.
References
-
- Abele R. and Tampé,R. (1999) Function of the transport complex TAP in cellular immune recognition. Biochim. Biophys. Acta, 1461, 405–419. - PubMed
-
- Abrahams J.P. and Leslie,A.G.W. (1996) Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr. D, 52, 30–42. - PubMed
-
- Alberts P., Daumke,O., Deverson,E.V., Howard,J.C. and Knittler,M.R. (2001) Distinct functional properties of the TAP subunits coordinate the nucleotide-dependent transport cycle. Curr. Biol., 11, 242–251. - PubMed
-
- Androlewicz M.J., Anderson,K.S. and Cresswell,P. (1993) Evidence that transporters associated with antigen processing translocate a major histocompatibility complex class I-binding peptide into the endoplasmic reticulum in an ATP-dependent manner. Proc. Natl Acad. Sci. USA, 90, 9130–9134. - PMC - PubMed
-
- Brünger A.T. et al. (1998) Crystallography and NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D, 54, 905–921. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

