Role of Agrobacterium VirB11 ATPase in T-pilus assembly and substrate selection
- PMID: 11566978
- PMCID: PMC99657
- DOI: 10.1128/JB.183.20.5813-5825.2001
Role of Agrobacterium VirB11 ATPase in T-pilus assembly and substrate selection
Abstract
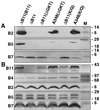
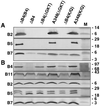
The VirB11 ATPase is a subunit of the Agrobacterium tumefaciens transfer DNA (T-DNA) transfer system, a type IV secretion pathway required for delivery of T-DNA and effector proteins to plant cells during infection. In this study, we examined the effects of virB11 mutations on VirB protein accumulation, T-pilus production, and substrate translocation. Strains synthesizing VirB11 derivatives with mutations in the nucleoside triphosphate binding site (Walker A motif) accumulated wild-type levels of VirB proteins but failed to produce the T-pilus or export substrates at detectable levels, establishing the importance of nucleoside triphosphate binding or hydrolysis for T-pilus biogenesis. Similar findings were obtained for VirB4, a second ATPase of this transfer system. Analyses of strains expressing virB11 dominant alleles in general showed that T-pilus production is correlated with substrate translocation. Notably, strains expressing dominant alleles previously designated class II (dominant and nonfunctional) neither transferred T-DNA nor elaborated detectable levels of the T-pilus. By contrast, strains expressing most dominant alleles designated class III (dominant and functional) efficiently translocated T-DNA and synthesized abundant levels of T pilus. We did, however, identify four types of virB11 mutations or strain genotypes that selectively disrupted substrate translocation or T-pilus production: (i) virB11/virB11* merodiploid strains expressing all class II and III dominant alleles were strongly suppressed for T-DNA translocation but efficiently mobilized an IncQ plasmid to agrobacterial recipients and also elaborated abundant levels of T pilus; (ii) strains synthesizing two class III mutant proteins, VirB11, V258G and VirB11.I265T, efficiently transferred both DNA substrates but produced low and undetectable levels of T pilus, respectively; (iii) a strain synthesizing the class II mutant protein VirB11.I103T/M301L efficiently exported VirE2 but produced undetectable levels of T pilus; (iv) strains synthesizing three VirB11 derivatives with a four-residue (HMVD) insertion (L75.i4, C168.i4, and L302.i4) neither transferred T-DNA nor produced detectable levels of T pilus but efficiently transferred VirE2 to plants and the IncQ plasmid to agrobacterial recipient cells. Together, our findings support a model in which the VirB11 ATPase contributes at two levels to type IV secretion, T-pilus morphogenesis, and substrate selection. Furthermore, the contributions of VirB11 to machine assembly and substrate transfer can be uncoupled by mutagenesis.
Figures





References
-
- Beijersbergen A, Smith S J, Hooykas P J J. Localization and topology of VirB proteins of Agrobacterium tumefaciens. Plasmid. 1994;32:212–218. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

