Methylation of the nucleobases in RNA oligonucleotides mediates duplex-hairpin conversion
- PMID: 11574682
- PMCID: PMC115353
- DOI: 10.1093/nar/29.19.3997
Methylation of the nucleobases in RNA oligonucleotides mediates duplex-hairpin conversion
Abstract
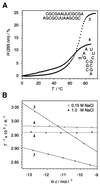
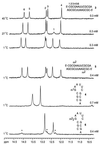
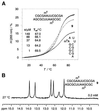
We have systematically investigated the duplex to hairpin conversion of oligoribonucleotides under the aspect of nucleobase methylation. The first part of our study refers to the self-complementary sequence rCGCGAAUUCGCGA, which forms a stable Watson-Crick base paired duplex under various buffer conditions. It is shown that this sequence is forced to adopt a hairpin conformation if one of the central 6 nt is replaced by the corresponding methylated nucleotide, such as 1-methylguanosine N(2),N(2)-dimethylguanosine, N(6),N(6)-dimethyladenosine (m(6)(2)A) or 3-methyluridine. On the other hand, the duplex structure is retained and even stabilized by replacement of a central nucleotide with N(2)-methylguanosine (m(2)G) or N(4)-methylcytidine. A borderline case is represented by N(6)-methyladenosine (m(6)A). Although generally a duplex-preserving modification, our data indicate that m(6)A in specific strand positions and at low strand concentrations is able to effectuate duplex-hairpin conversion. Our studies also include the ssu ribosomal helix 45 sequence motif, rGACCm(2)GGm(6)(2)Am(6)(2)AGGUC. In analogy, it is demonstrated that the tandem m(6)(2)A nucleobases of this oligoribonucleotide prevent duplex formation with complementary strands. Therefore, it can be concluded that nucleobase methylations at the Watson-Crick base pairing site provide the potential not only to modulate but to substantially affect RNA structure by formation of different secondary structure motifs.
Figures



References
-
- Björk G.R. (1995) Biosynthesis and function of modified nucleosides. In Söll,D. and RajBhandary,U. (eds), tRNA: Structure, Biosynthesis and Function. ASM Press, Washington, DC, pp. 165–205.
-
- Yokoyama S. and Nishimura,S. (1995) Modified nucleosides and codon recognition. In Söll,D. and RajBhandary,U. (eds), tRNA: Structure, Biosynthesis and Function. ASM Press, Washington, DC, pp. 207–223.
-
- Motorin Y. and Grosjean,H. (1998) Appendix 1: Chemical structures and classification of post-transcriptionally modified nucleosides in RNA. In Grosjean,H. and Benne,R. (eds), Modification and Editing of RNA. ASM Press, Washington, DC, pp. 543–549.
-
- Grosjean H. and Benne,R. (eds) (1998) Modification and Editing of RNA. ASM Press, Washington, DC.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

