Absence of Dbp2p alters both nonsense-mediated mRNA decay and rRNA processing
- PMID: 11585918
- PMCID: PMC99910
- DOI: 10.1128/MCB.21.21.7366-7379.2001
Absence of Dbp2p alters both nonsense-mediated mRNA decay and rRNA processing
Abstract
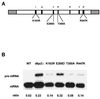
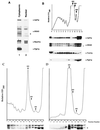
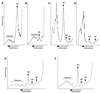
Dbp2p, a member of the large family of DEAD-box proteins and a yeast homolog of human p68, was shown to interact with Upf1p, an essential component of the nonsense-mediated mRNA decay pathway. Dbp2p:Upf1p interaction occurs within a large conserved region in the middle of Upf1p that is largely distinct from its Nmd2p and Sup35/45p interaction domains. Deletion of DBP2, or point mutations within its highly conserved DEAD-box motifs, increased the abundance of nonsense-containing transcripts, leading us to conclude that Dbp2p also functions in the nonsense-mediated mRNA decay pathway. Dbp2p, like Upf1p, acts before or at decapping, is predominantly cytoplasmic, and associates with polyribosomes. Interestingly, Dbp2p also plays an important role in rRNA processing. In dbp2Delta cells, polyribosome profiles are deficient in free 60S subunits and the mature 25S rRNA is greatly reduced. The ribosome biogenesis phenotype, but not the mRNA decay function, of dbp2Delta cells can be complemented by the human p68 gene. We propose a unifying model in which Dbp2p affects both nonsense-mediated mRNA decay and rRNA processing by altering rRNA structure, allowing specific processing events in one instance and facilitating dissociation of the translation termination complex in the other.
Figures






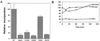



Similar articles
-
Identification of a novel component of the nonsense-mediated mRNA decay pathway by use of an interacting protein screen.Genes Dev. 1995 Feb 15;9(4):437-54. doi: 10.1101/gad.9.4.437. Genes Dev. 1995. PMID: 7883168
-
Upf1p, Nmd2p, and Upf3p regulate the decapping and exonucleolytic degradation of both nonsense-containing mRNAs and wild-type mRNAs.Mol Cell Biol. 2001 Mar;21(5):1515-30. doi: 10.1128/MCB.21.5.1515-1530.2001. Mol Cell Biol. 2001. PMID: 11238889 Free PMC article.
-
Upf1p, a highly conserved protein required for nonsense-mediated mRNA decay, interacts with the nuclear pore proteins Nup100p and Nup116p.Gene. 2005 Jan 31;345(2):199-212. doi: 10.1016/j.gene.2004.10.005. Epub 2004 Dec 15. Gene. 2005. PMID: 15716093
-
[Role and mechanism of NMD (nonsense-mediated mRNA decay)].Tanpakushitsu Kakusan Koso. 2003 Mar;48(4 Suppl):382-9. Tanpakushitsu Kakusan Koso. 2003. PMID: 12696145 Review. Japanese. No abstract available.
-
An 'Arms Race' between the Nonsense-mediated mRNA Decay Pathway and Viral Infections.Semin Cell Dev Biol. 2021 Mar;111:101-107. doi: 10.1016/j.semcdb.2020.05.018. Epub 2020 Jun 15. Semin Cell Dev Biol. 2021. PMID: 32553580 Free PMC article. Review.
Cited by
-
The RNA helicase Ddx5/p68 binds to hUpf3 and enhances NMD of Ddx17/p72 and Smg5 mRNA.Nucleic Acids Res. 2013 Sep;41(16):7875-88. doi: 10.1093/nar/gkt538. Epub 2013 Jun 20. Nucleic Acids Res. 2013. PMID: 23788676 Free PMC article.
-
The DEAD-box protein Dbp2p is linked to noncoding RNAs, the helicase Sen1p, and R-loops.RNA. 2018 Dec;24(12):1693-1705. doi: 10.1261/rna.067249.118. Epub 2018 Sep 27. RNA. 2018. PMID: 30262458 Free PMC article.
-
The putative RNA helicase Dbp6p functionally interacts with Rpl3p, Nop8p and the novel trans-acting Factor Rsa3p during biogenesis of 60S ribosomal subunits in Saccharomyces cerevisiae.Genetics. 2004 Apr;166(4):1687-99. doi: 10.1534/genetics.166.4.1687. Genetics. 2004. PMID: 15126390 Free PMC article.
-
P68 RNA helicase as a molecular target for cancer therapy.J Exp Clin Cancer Res. 2014 Aug 24;33(1):64. doi: 10.1186/s13046-014-0064-y. J Exp Clin Cancer Res. 2014. PMID: 25150365 Free PMC article. Review.
-
Comprehensive analysis of the Kinetoplastea intron landscape reveals a novel intron-containing gene and the first exclusively trans-splicing eukaryote.BMC Biol. 2024 Dec 3;22(1):281. doi: 10.1186/s12915-024-02080-z. BMC Biol. 2024. PMID: 39627879 Free PMC article.
References
-
- Atkin A L, Schenkman L R, Eastman M, Dahlseid J N, Lelivelt M J, Culbertson M R. Relationship between yeast polyribosomes and Upf proteins required for nonsense mRNA decay. J Biol Chem. 1997;272:22163–22172. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
