Cytoplasmic dynein light intermediate chain is required for discrete aspects of mitosis in Caenorhabditis elegans
- PMID: 11598181
- PMCID: PMC60145
- DOI: 10.1091/mbc.12.10.2921
Cytoplasmic dynein light intermediate chain is required for discrete aspects of mitosis in Caenorhabditis elegans
Abstract
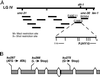
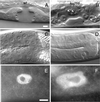
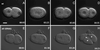
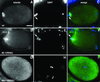
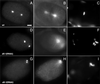
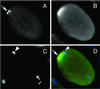
We describe phenotypic characterization of dli-1, the Caenorhabditis elegans homolog of cytoplasmic dynein light intermediate chain (LIC), a subunit of the cytoplasmic dynein motor complex. Animals homozygous for loss-of-function mutations in dli-1 exhibit stochastic failed divisions in late larval cell lineages, resulting in zygotic sterility. dli-1 is required for dynein function during mitosis. Depletion of the dli-1 gene product through RNA-mediated gene interference (RNAi) reveals an early embryonic requirement. One-cell dli-1(RNAi) embryos exhibit failed cell division attempts, resulting from a variety of mitotic defects. Specifically, pronuclear migration, centrosome separation, and centrosome association with the male pronuclear envelope are defective in dli-1(RNAi) embryos. Meiotic spindle formation, however, is not affected in these embryos. DLI-1, like its vertebrate homologs, contains a putative nucleotide-binding domain similar to those found in the ATP-binding cassette transporter family of ATPases as well as other nucleotide-binding and -hydrolyzing proteins. Amino acid substitutions in a conserved lysine residue, known to be required for nucleotide binding, confers complete rescue in a dli-1 mutant background, indicating this is not an essential domain for DLI-1 function.
Figures


Similar articles
-
Cytoplasmic dynein is required for distinct aspects of MTOC positioning, including centrosome separation, in the one cell stage Caenorhabditis elegans embryo.J Cell Biol. 1999 Oct 4;147(1):135-50. doi: 10.1083/jcb.147.1.135. J Cell Biol. 1999. PMID: 10508861 Free PMC article.
-
Functional analysis of cytoplasmic dynein heavy chain in Caenorhabditis elegans with fast-acting temperature-sensitive mutations.Mol Biol Cell. 2005 Mar;16(3):1200-12. doi: 10.1091/mbc.e04-06-0523. Epub 2004 Dec 22. Mol Biol Cell. 2005. PMID: 15616192 Free PMC article.
-
Computer simulations reveal mechanisms that organize nuclear dynein forces to separate centrosomes.Mol Biol Cell. 2017 Nov 7;28(23):3165-3170. doi: 10.1091/mbc.E16-12-0823. Epub 2017 Jul 12. Mol Biol Cell. 2017. PMID: 28701341 Free PMC article.
-
What can Caenorhabditis elegans tell us about the nuclear envelope?FEBS Lett. 2007 Jun 19;581(15):2794-801. doi: 10.1016/j.febslet.2007.03.052. Epub 2007 Mar 30. FEBS Lett. 2007. PMID: 17418822 Review.
-
The elegans of spindle assembly.Cell Mol Life Sci. 2010 Jul;67(13):2195-213. doi: 10.1007/s00018-010-0324-8. Epub 2010 Mar 26. Cell Mol Life Sci. 2010. PMID: 20339898 Free PMC article. Review.
Cited by
-
Polarized endosome dynamics engage cytoplasmic Par-3 that recruits dynein during asymmetric cell division.Sci Adv. 2021 Jun 11;7(24):eabg1244. doi: 10.1126/sciadv.abg1244. Print 2021 Jun. Sci Adv. 2021. PMID: 34117063 Free PMC article.
-
UNC-16/JIP3 and UNC-76/FEZ1 limit the density of mitochondria in C. elegans neurons by maintaining the balance of anterograde and retrograde mitochondrial transport.Sci Rep. 2018 Jun 12;8(1):8938. doi: 10.1038/s41598-018-27211-9. Sci Rep. 2018. PMID: 29895958 Free PMC article.
-
Recruitment of dynein to late endosomes and lysosomes through light intermediate chains.Mol Biol Cell. 2011 Feb 15;22(4):467-77. doi: 10.1091/mbc.E10-02-0129. Epub 2010 Dec 17. Mol Biol Cell. 2011. PMID: 21169557 Free PMC article.
-
A ZYG-12-dynein interaction at the nuclear envelope defines cytoskeletal architecture in the C. elegans gonad.J Cell Biol. 2009 Jul 27;186(2):229-41. doi: 10.1083/jcb.200902101. J Cell Biol. 2009. PMID: 19635841 Free PMC article.
-
Uncovering buffered pleiotropy: a genome-scale screen for mel-28 genetic interactors in Caenorhabditis elegans.G3 (Bethesda). 2014 Jan 10;4(1):185-96. doi: 10.1534/g3.113.008532. G3 (Bethesda). 2014. PMID: 24281427 Free PMC article.
References
-
- Albertson DG. C. elegans II. In: Riddle DL, Blumenthal T, Meyer BJ, Priess JR, editors. C. elegans II. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 1997. pp. 47–78.
-
- Barstead RJ, Kleiman L, Waterston RH. Cloning, sequencing, and mapping of an alpha-actinin gene from the nematode Caenorhabditis elegans. Cell Motil Cytoskeleton. 1991;20:69–78. - PubMed
-
- Bielli A, Thornqvist PO, Hendrick AG, Finn R, Fitzgerald K, McCaffrey MW. The small GTPase Rab4A interacts with the central region of cytoplasmic dynein light intermediate chain-1. Biochem Biophys Res Commun. 2001;281:1141–1153. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

