SUMO modification of Rad22, the Schizosaccharomyces pombe homologue of the recombination protein Rad52
- PMID: 11600706
- PMCID: PMC60211
- DOI: 10.1093/nar/29.20.4179
SUMO modification of Rad22, the Schizosaccharomyces pombe homologue of the recombination protein Rad52
Abstract
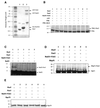
The Schizosaccharomyces pombe rad31 and hus5 genes are required for the DNA damage response, as mutants defective in these genes are sensitive to DNA damaging agents, such as UV and ionising radiation and to the DNA synthesis inhibitor hydroxyurea (HU). Sequence analysis has suggested that rad31 and hus5 encode components of the Pmt3 (SUMO) modification process in S.pombe. We show here that the rad31 null and hus5.62 mutants display reduced levels of Pmt3 modification. We have initiated a search for proteins required for the DNA damage response, which may be modified by Pmt3 and have identified Rad22, the fission yeast homologue of the recombination protein Rad52. Purification of myc + His-tagged Rad22 protein from cells expressing HA-tagged Pmt3 identifies an 83 kDa species which cross-reacts with anti-HA antisera. We show here that Rad22 interacts with Rhp51 and Rpa70 (the fission yeast homologues of Rad51 and the large subunit of RPA, respectively), but that neither of these proteins appears to be responsible for the 83 kDa species. The 83 kDa species is observed when extracts are prepared under both native and denaturing conditions, and is also observed when myc + His-tagged Rad22 and Pmt3 are expressed at wild type levels, suggesting that Rad22 is modified by Pmt3 in vivo. We have established an S.pombe in vitro Pmt3 modification system and have shown that Rad22 and Rhp51 are modified in vitro, but that Rpa70 is not.
Figures



Similar articles
-
Characterization of SUMO-conjugating enzyme mutants in Schizosaccharomyces pombe identifies a dominant-negative allele that severely reduces SUMO conjugation.Biochem J. 2003 May 15;372(Pt 1):97-104. doi: 10.1042/BJ20021645. Biochem J. 2003. PMID: 12597774 Free PMC article.
-
Physical interaction between recombinational proteins Rhp51 and Rad22 in Schizosaccharomyces pombe.J Biol Chem. 2002 Aug 16;277(33):30264-70. doi: 10.1074/jbc.M202517200. Epub 2002 Jun 5. J Biol Chem. 2002. PMID: 12050150
-
Cell-cycle-dependent localisation of Ulp1, a Schizosaccharomyces pombe Pmt3 (SUMO)-specific protease.J Cell Sci. 2002 Mar 15;115(Pt 6):1113-22. doi: 10.1242/jcs.115.6.1113. J Cell Sci. 2002. PMID: 11884512
-
Molecular biology of DNA repair in the fission yeast Schizosaccharomyces pombe.Mutat Res. 1996 Aug 8;363(3):147-61. doi: 10.1016/0921-8777(96)00017-1. Mutat Res. 1996. PMID: 8765156 Review. No abstract available.
-
Repair of UV damage in the fission yeast Schizosaccharomyces pombe.Mutat Res. 2000 Jun 30;451(1-2):197-210. doi: 10.1016/s0027-5107(00)00050-6. Mutat Res. 2000. PMID: 10915873 Review.
Cited by
-
Role of the fission yeast SUMO E3 ligase Pli1p in centromere and telomere maintenance.EMBO J. 2004 Oct 1;23(19):3844-53. doi: 10.1038/sj.emboj.7600394. Epub 2004 Sep 9. EMBO J. 2004. PMID: 15359282 Free PMC article.
-
Genetic analysis of SUMOylation in Arabidopsis: conjugation of SUMO1 and SUMO2 to nuclear proteins is essential.Plant Physiol. 2007 Sep;145(1):119-34. doi: 10.1104/pp.107.102285. Epub 2007 Jul 20. Plant Physiol. 2007. PMID: 17644626 Free PMC article.
-
Fission yeast Rad52 phosphorylation restrains error prone recombination pathways.PLoS One. 2014 Apr 18;9(4):e95788. doi: 10.1371/journal.pone.0095788. eCollection 2014. PLoS One. 2014. PMID: 24748152 Free PMC article.
-
The C-terminal zinc finger of the catalytic subunit of DNA polymerase delta is responsible for direct interaction with the B-subunit.Nucleic Acids Res. 2004 Jun 1;32(10):3005-16. doi: 10.1093/nar/gkh623. Print 2004. Nucleic Acids Res. 2004. PMID: 15173383 Free PMC article.
-
SUMOylation is required for normal development of linear elements and wild-type meiotic recombination in Schizosaccharomyces pombe.Chromosoma. 2010 Feb;119(1):59-72. doi: 10.1007/s00412-009-0241-5. Epub 2009 Sep 12. Chromosoma. 2010. PMID: 19756689
References
-
- Taylor E.M. and Lehmann,A.R. (1998) Conservation of eukaryotic DNA repair mechanisms. Int. J. Radiat. Biol., 74, 277–286. - PubMed
-
- Caspari T. and Carr,A.M. (1999) DNA structure checkpoint pathways in Schizosaccharomyces pombe. Biochimie, 81, 173–181. - PubMed
-
- al-Khodairy F., Enoch,T., Hagan,I.M. and Carr,A.M. (1995) The Schizosaccharomyces pombe hus5 gene encodes a ubiquitin conjugating enzyme required for normal mitosis. J. Cell Sci., 108, 475–486. - PubMed
-
- Johnson E.S. and Blobel,G. (1997) Ubc9p is the conjugating enzyme for the ubiquitin-like protein Smt3p. J. Biol. Chem., 272, 26799–26802. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

