Transgenically expressed T-Rep of tomato yellow leaf curl Sardinia virus acts as a trans-dominant-negative mutant, inhibiting viral transcription and replication
- PMID: 11602699
- PMCID: PMC114639
- DOI: 10.1128/JVI.75.22.10573-10581.2001
Transgenically expressed T-Rep of tomato yellow leaf curl Sardinia virus acts as a trans-dominant-negative mutant, inhibiting viral transcription and replication
Abstract
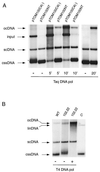
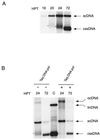
We have previously shown that transgenic expression of a truncated C1 gene of Tomato yellow leaf curl Sardinia virus (TYLCSV), expressing the first 210 amino acids of the replication-associated protein (T-Rep) and potentially coexpressing the C4 protein, confers resistance to the homologous virus in Nicotiana benthamiana plants. In the present study we have investigated the role of T-Rep and C4 proteins in the resistance mechanism, analyzing changes in virus transcription and replication. Transgenic plants and protoplasts were challenged with TYLCSV and the related TYLCSV Murcia strain (TYLCSV-ES[1]). TYLCSV-resistant plants were susceptible to TYLCSV-ES[1]; moreover, TYLCSV but not TYLCSV-ES[1] replication was strongly inhibited in transgenic protoplasts as well as in wild-type (wt) protoplasts transiently expressing T-Rep but not the C4 protein. Viral circular single-stranded DNA (cssDNA) was usually undetectable in transgenically and transiently T-Rep-expressing protoplasts, while viral DNAs migrating more slowly than the cssDNA were observed. Biochemical studies showed that these DNAs were partial duplexes with the minus strand incomplete. Interestingly, similar viral DNA forms were also found at early stages of TYLCSV replication in wt N. benthamiana protoplasts. Transgenically expressed T-Rep repressed the transcription of the GUS reporter gene up to 300-fold when fused to the homologous (TYLCSV) but not to the heterologous (TYLCSV-ES[1]) C1 promoter. Similarly, transiently expressed T-Rep but not C4 protein strongly repressed GUS transcription when fused to the C1 promoter of TYLCSV. A model of T-Rep interference with TYLCSV transcription-replication is proposed.
Figures




References
-
- Bouche J P, Rowen L, Kornberg A. The RNA primer synthesized by primase to initiate phage G4 DNA replication. J Biol Chem. 1978;253:765–769. - PubMed
-
- Brunetti A, Tavazza M, Noris E, Tavazza R, Caciagli P, Ancora G, Crespi S, Accotto G P. High expression of truncated viral Rep protein confers resistance to tomato yellow leaf curl virus in transgenic tomato plants. Mol Plant-Microbe Interact. 1997;10:571–579.
-
- Chatterji A, Chatterji U, Beachy R N, Fauquet C M. Sequence parameters that determine specificity of binding of the replication-associated protein to its cognate site in two strains of tomato leaf curl virus-New Delhi. Virology. 2000;273:341–350. - PubMed
-
- Choi I R, Stenger D C. Strain-specific determinants of beet curly top geminivirus DNA replication. Virology. 1995;206:904–912. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

