On the molecular discrimination between adenine and guanine by proteins
- PMID: 11691917
- PMCID: PMC60199
- DOI: 10.1093/nar/29.21.4294
On the molecular discrimination between adenine and guanine by proteins
Abstract
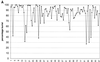
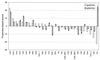
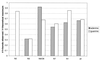
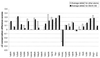
The molecular recognition and discrimination of adenine and guanine ligand moieties in complexes with proteins have been studied using empirical observations on carefully selected crystal structures. The distribution of protein folds that bind these purines has been found to differ significantly from that across the whole PDB, but the most populated architectures and folds are also the most common in three genomes from the three different domains of life. The protein environments around the two nucleic acid bases were significantly different, in terms of the propensities of amino acid residues to be in the binding site, as well as their propensities to form hydrogen bonds to the bases. Plots of the distribution of protein atoms around the two purines clearly show different clustering of hydrogen bond donors and acceptors opposite complimentary acceptors and donors in the rings, with hydrophobic areas below and above the rings. However, the clustering pattern is fuzzy, reflecting the variety of ways that proteins have evolved to recognise the same molecular moiety. Furthermore, an analysis of the conservation of residues in the protein chains binding guanine shows that residues in contact with the base are in general better conserved than the rest of the chain.
Figures
















References
-
- Lommerse J.P.M., Price,S.L. and Taylor,R. (1997) Hydrogen bonding of carbonyl, ether, and ester oxygen atoms with alkanol hydroxyl groups. J. Comput. Chem., 18, 757–774.
-
- Nobeli I., Price,S.L., Lommerse,J.P.M. and Taylor,R. (1997) Hydrogen bonding properties of oxygen and nitrogen acceptors in aromatic heterocycles. J. Comput. Chem., 18, 2060–2074.
-
- Dunitz J.D. and Taylor,R. (1997) Organic fluorine hardly ever accepts hydrogen bonds. Chem. Eur. J., 3, 89–98.
-
- Allen F.H., Baalham,C.A., Lommerse,J.P.M., Raithby,P.R. and Sparr,E. (1997) Hydrogen-bond acceptor properties of nitro-O atoms: A combined crystallographic database and ab initio molecular orbital study. Acta Crystallogr., B53, 1017–1024.
-
- Bruno I.J., Cole,J.C., Lommerse,J.P.M., Rowland,R.S., Taylor,R. and Verdonk,M.L. (1997) IsoStar: a library of information about nonbonded interactions. J. Comput. Aid. Mol. Des., 11, 525–537. - PubMed

