Inhibition of CHOP translation by a peptide encoded by an open reading frame localized in the chop 5'UTR
- PMID: 11691921
- PMCID: PMC60176
- DOI: 10.1093/nar/29.21.4341
Inhibition of CHOP translation by a peptide encoded by an open reading frame localized in the chop 5'UTR
Abstract
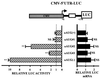
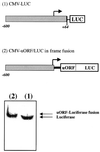
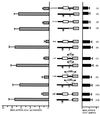
Chop is a ubiquitously expressed mammalian gene encoding a small nuclear protein related to the CCAAT/enhancer-binding protein (C/EBP) family of transcription factors. CHOP protein plays an important role in various cellular processes such as growth, differentiation and programmed cell death. CHOP expression is strongly increased in response to a large variety of stresses including perturbation of the endoplasmic reticulum function, DNA damage and nutrient deprivation. Multiple mechanisms including transcriptional and post-transcriptional controls are involved in the regulation of CHOP expression. We show here that the 5'UTR of the Chop transcript plays an important role in controlling the synthesis of CHOP protein. In particular, the 5'UTR contains a conserved uORF which encodes a 31 amino acid peptide that inhibits the expression of the downstream ORF. Mutational analysis of the 5' leader region and peptide coding sequences suggests that the peptide itself inhibits expression of the downstream ORF. Such results suggest a role for uORF in limiting ribosomal access to downstream initiation sites. With respect to the importance of CHOP protein in the regulation of cellular functions, the mechanisms that regulate its basal level are of considerable interest.
Figures



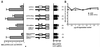
Similar articles
-
Analysis of full length ADAMTS6 transcript reveals alternative splicing and a role for the 5' untranslated region in translational control.Gene. 2005 Oct 10;359:99-110. doi: 10.1016/j.gene.2005.06.011. Gene. 2005. PMID: 16129570
-
Efficient translation initiation directed by the 900-nucleotide-long and GC-rich 5' untranslated region of the human retrotransposon LINE-1 mRNA is strictly cap dependent rather than internal ribosome entry site mediated.Mol Cell Biol. 2007 Jul;27(13):4685-97. doi: 10.1128/MCB.02138-06. Epub 2007 Apr 30. Mol Cell Biol. 2007. PMID: 17470553 Free PMC article.
-
Post-termination ribosome interactions with the 5'UTR modulate yeast mRNA stability.EMBO J. 1999 Jun 1;18(11):3139-52. doi: 10.1093/emboj/18.11.3139. EMBO J. 1999. PMID: 10357825 Free PMC article.
-
Translational control by 5'-untranslated regions of eukaryotic mRNAs.Science. 2016 Jun 17;352(6292):1413-6. doi: 10.1126/science.aad9868. Science. 2016. PMID: 27313038 Free PMC article. Review.
-
The interplay between cis- and trans-acting factors drives selective mRNA translation initiation in eukaryotes.Biochimie. 2024 Feb;217:20-30. doi: 10.1016/j.biochi.2023.09.017. Epub 2023 Sep 21. Biochimie. 2024. PMID: 37741547 Review.
Cited by
-
Genome-wide search for novel human uORFs and N-terminal protein extensions using ribosomal footprinting.Genome Res. 2012 Nov;22(11):2208-18. doi: 10.1101/gr.139568.112. Epub 2012 Aug 9. Genome Res. 2012. PMID: 22879431 Free PMC article.
-
Peptides encoded by noncoding genes: challenges and perspectives.Signal Transduct Target Ther. 2019 Dec 13;4:57. doi: 10.1038/s41392-019-0092-3. eCollection 2019. Signal Transduct Target Ther. 2019. PMID: 31871775 Free PMC article. Review.
-
DDIT3 Expression in Liposarcoma Development.Sarcoma. 2014;2014:954671. doi: 10.1155/2014/954671. Epub 2014 Mar 25. Sarcoma. 2014. PMID: 24790523 Free PMC article.
-
Ribosome Elongation Stall Directs Gene-specific Translation in the Integrated Stress Response.J Biol Chem. 2016 Mar 18;291(12):6546-58. doi: 10.1074/jbc.M115.705640. Epub 2016 Jan 27. J Biol Chem. 2016. PMID: 26817837 Free PMC article.
-
MCTS2 and distinct eIF2D roles in uORF-dependent translation regulation revealed by in vitro re-initiation assays.EMBO J. 2025 Feb;44(3):854-876. doi: 10.1038/s44318-024-00347-3. Epub 2025 Jan 2. EMBO J. 2025. PMID: 39748120 Free PMC article.
References
-
- McKnight S.L., Lane,M.D. and Gluecksohn-Waelsch,S. (1989) Is CCAAT/enhancer-binding protein a central regulator of energy metabolism? Genes Dev., 3, 2021–2024. - PubMed
-
- Umek R.M., Friedman,A.D. and McKnight,S.L. (1991) CCAAT-enhancer binding protein: a component of a differentiation switch. Science, 251, 288–292. - PubMed
-
- Cao Z., Umek,R.M. and McKnight,S.L. (1991) Regulated expression of three C/EBP isoforms during adipose conversion of 3T3-L1 cells. Genes Dev., 5, 1538–1552. - PubMed
-
- Birkenmeier E.H., Gwynn,B., Howard,S., Jerry,J., Gordon,J.I., Landschulz,W.H. and McKnight,S.L. (1989) Tissue-specific expression, developmental regulation and genetic mapping of the gene encoding CCAAT/enhancer binding protein. Genes Dev., 3, 1146–1156. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

