IS186 insertion at a hot spot in the lon promoter as a basis for lon protease deficiency of Escherichia coli B: identification of a consensus target sequence for IS186 transposition
- PMID: 11698384
- PMCID: PMC95536
- DOI: 10.1128/JB.183.23.6943-6946.2001
IS186 insertion at a hot spot in the lon promoter as a basis for lon protease deficiency of Escherichia coli B: identification of a consensus target sequence for IS186 transposition
Abstract
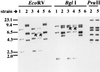
The radiation sensitivity of Escherichia coli B was first described more than 50 years ago, and the genetic locus responsible for the trait was subsequently identified as lon (encoding Lon protease). We now show that both E. coli B and the first reported E. coli K-12 lon mutant, AB1899, carry IS186 insertions in opposite orientations at a single site in the lon promoter region and that this site represents a natural hot spot for transposition of the insertion sequence (IS) element. Our analysis of deposited sequence data for a number of other IS186 insertion sites permitted the deductions that (i) the consensus target site sequence for IS186 transposition is 5'-(G)(> or =4)(N)(3-6)(C)(> or =4)-3', (ii) the associated host sequence duplication varies within the range of 6 to 12 bp and encompasses the N(3-6) sequence, and (iii) in a majority of instances, at least one end of the duplication is at the G-N (or N-C) junction. IS186-related sequences were absent in closely related bacterium Salmonella enterica serovar Typhimurium, indicating that this IS element is a recent acquisition in the evolutionary history of E. coli.
Figures

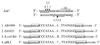
Similar articles
-
Cloning, expression, and characterization of the lon gene of Erwinia amylovora: evidence for a heat shock response.J Bacteriol. 1995 Feb;177(4):932-7. doi: 10.1128/jb.177.4.932-937.1995. J Bacteriol. 1995. PMID: 7860603 Free PMC article.
-
[Integration of IS186 element into the -10-region of the promoter of heat shock lon-gene in Escherichia coli].Bioorg Khim. 1997 Jan;23(1):18-20. Bioorg Khim. 1997. PMID: 9139638 Russian.
-
Molecular cloning of the Lon protease gene from Thermus thermophilus HB8 and characterization of its gene product.Eur J Biochem. 1999 Dec;266(3):811-9. doi: 10.1046/j.1432-1327.1999.00907.x. Eur J Biochem. 1999. PMID: 10583374
-
Sequence of the lon gene in Escherichia coli. A heat-shock gene which encodes the ATP-dependent protease La.J Biol Chem. 1988 Aug 25;263(24):11718-28. J Biol Chem. 1988. PMID: 3042779
-
[Structural and functional characteristics of ATP-dependent Lon-proteinase from Escherichia coli].Bioorg Khim. 1999 Dec;25(12):883-91. Bioorg Khim. 1999. PMID: 10734549 Review. Russian.
Cited by
-
Genome rearrangements in Escherichia coli during de novo acquisition of resistance to a single antibiotic or two antibiotics successively.BMC Genomics. 2018 Dec 27;19(1):973. doi: 10.1186/s12864-018-5353-y. BMC Genomics. 2018. PMID: 30591014 Free PMC article.
-
Increased genome instability in Escherichia coli lon mutants: relation to emergence of multiple-antibiotic-resistant (Mar) mutants caused by insertion sequence elements and large tandem genomic amplifications.Antimicrob Agents Chemother. 2007 Apr;51(4):1293-303. doi: 10.1128/AAC.01128-06. Epub 2007 Jan 12. Antimicrob Agents Chemother. 2007. PMID: 17220404 Free PMC article.
-
Altered expression of a quality control protease in E. coli reshapes the in vivo mutational landscape of a model enzyme.Elife. 2020 Jul 23;9:e53476. doi: 10.7554/eLife.53476. Elife. 2020. PMID: 32701056 Free PMC article.
-
The quorum-sensing protein TraR of Agrobacterium tumefaciens is susceptible to intrinsic and TraM-mediated proteolytic instability.Mol Microbiol. 2012 Jun;84(5):807-15. doi: 10.1111/j.1365-2958.2012.08037.x. Epub 2012 Apr 19. Mol Microbiol. 2012. PMID: 22515735 Free PMC article.
-
Comparative multi-omics systems analysis of Escherichia coli strains B and K-12.Genome Biol. 2012 May 25;13(5):R37. doi: 10.1186/gb-2012-13-5-r37. Genome Biol. 2012. PMID: 22632713 Free PMC article.
References
-
- Agrawal A, Eastman Q M, Schatz D G. Transposition mediated by RAG1 and RAG2 and its implications for the evolution of the immune system. Nature. 1998;394:744–751. - PubMed
-
- Birkenbihl R P, Vielmetter W. Completion of the IS map in E. coli: IS186 positions on the E. coli K12 chromosome. Mol Gen Genet. 1991;226:318–320. - PubMed
-
- Blattner F R, Plunkett III G, Bloch C A, Perna N T, Burland V, Riley M, Collado-Vides J, Glasner J D, Rode C K, Mayhew G F, Gregor J, Davis N W, Kirkpatrick H A, Goeden M A, Rose D J, Mau B, Shao Y. The complete genome sequence of Escherichia coli K-12. Science. 1997;277:1453–1462. - PubMed
-
- Chong P, Hui I, Loo T, Gillam S. Structural analysis of a new GC-specific insertion element IS186. FEBS Lett. 1985;192:47–52. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

