Phase variation in the Helicobacter pylori phospholipase A gene and its role in acid adaptation
- PMID: 11705905
- PMCID: PMC98819
- DOI: 10.1128/IAI.69.12.7334-7340.2001
Phase variation in the Helicobacter pylori phospholipase A gene and its role in acid adaptation
Abstract
Previously, we have shown that Helicobacter pylori can spontaneously and reversibly change its membrane lipid composition, producing variants with low or high content of lysophospholipids. The "lyso" variant contains a high percentage of lysophospholipids, adheres better to epithelial cells, and releases more proteins such as urease and VacA, compared to the "normal" variant, which has a low content of lysophospholipids. Prolonged growth of the normal variant at pH 3.5, but not under neutral conditions, leads to enrichment of lyso variant colonies, suggesting that the colony switch is relevant to acid adaptation. In this study we show that the change in membrane lipid composition is due to phase variation in the pldA gene. A change in the (C) tract length of this gene results in reversible frameshifts, translation of a full-length or truncated pldA, and the production of active or inactive outer membrane phospholipase A (OMPLA). The role of OMPLA in determining the colony morphology was confirmed by the construction of an OMPLA-negative mutant. Furthermore, variants with an active OMPLA were able to survive acidic conditions better than variants with the inactive form. This explains why the lyso variant is selected at low pH. Our studies demonstrate that phase variation in the pldA gene, resulting in an active form of OMPLA, is important for survival under acidic conditions. We also demonstrated the active OMPLA genotype in fresh isolates of H. pylori from patients referred to gastroscopy for dyspepsia.
Figures
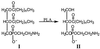




Similar articles
-
In silico evolutionary analysis of Helicobacter pylori outer membrane phospholipase A (OMPLA).BMC Microbiol. 2012 Sep 13;12:206. doi: 10.1186/1471-2180-12-206. BMC Microbiol. 2012. PMID: 22974200 Free PMC article.
-
High relative content of lysophospholipids of Helicobacter pylori mediates increased risk for ulcer disease.FEMS Immunol Med Microbiol. 2005 Apr 1;44(1):17-23. doi: 10.1016/j.femsim.2004.10.003. FEMS Immunol Med Microbiol. 2005. PMID: 15780574
-
Cholesteryl-6-O-acyl-alpha-D-glucopyranoside of Helicobacter pylori relate to relative lysophospholipid content.FEMS Microbiol Lett. 2005 Mar 1;244(1):117-20. doi: 10.1016/j.femsle.2005.01.030. FEMS Microbiol Lett. 2005. PMID: 15727830
-
Acid-responsive gene regulation in the human pathogen Helicobacter pylori.J Biotechnol. 2006 Oct 20;126(1):52-60. doi: 10.1016/j.jbiotec.2006.03.045. Epub 2006 May 19. J Biotechnol. 2006. PMID: 16713649 Review.
-
Outer-membrane phospholipase A: known structure, unknown biological function.Mol Microbiol. 2000 Feb;35(4):711-7. doi: 10.1046/j.1365-2958.2000.01775.x. Mol Microbiol. 2000. PMID: 10692149 Review.
Cited by
-
New implications on genomic adaptation derived from the Helicobacter pylori genome comparison.PLoS One. 2011 Feb 28;6(2):e17300. doi: 10.1371/journal.pone.0017300. PLoS One. 2011. PMID: 21387011 Free PMC article.
-
In silico evolutionary analysis of Helicobacter pylori outer membrane phospholipase A (OMPLA).BMC Microbiol. 2012 Sep 13;12:206. doi: 10.1186/1471-2180-12-206. BMC Microbiol. 2012. PMID: 22974200 Free PMC article.
-
Molecular mimicry in Helicobacter pylori infections.World J Gastroenterol. 2017 Jun 14;23(22):3964-3977. doi: 10.3748/wjg.v23.i22.3964. World J Gastroenterol. 2017. PMID: 28652651 Free PMC article. Review.
-
Outer membrane phospholipase A's roles in Helicobacter pylori acid adaptation.Gut Pathog. 2017 Jun 12;9:36. doi: 10.1186/s13099-017-0184-y. eCollection 2017. Gut Pathog. 2017. PMID: 28616083 Free PMC article.
-
PCR Based Detection of Phase Variable Genes in Pakistani Based Clinical Helicobacter pylori Strains.Jundishapur J Microbiol. 2016 Jul 3;9(7):e31824. doi: 10.5812/jjm.31824. eCollection 2016 Jul. Jundishapur J Microbiol. 2016. PMID: 27679701 Free PMC article.
References
-
- Alm R A, Ling L S, Moir D T, King B L, Brown E D, Doig P C, Smith D R, Noonan B, Guild B C, de Jonge B L, Carmel G, Tummino P J, Caruso A, Uria-Nickelsen M, Mills D M, Ives C, Gibson R, Merberg D, Mills S D, Jiang Q, Taylor D E, Vovis G F, Trust T J. Genomic-sequence comparison of two unrelated isolates of the human gastric pathogen Helicobacter pylori. Nature. 1999;397:176–180. - PubMed
-
- Appelmelk B J, Martin S L, Monteiro M, Clayton C A, McColm A A, Zheng P, Verboom T, Maaskant J J, van den Eijnden D H, Wirth H-P, Hokke C H, Perry M B, Vandenbroucke-Grauls C M J E, Kusters J G. Phase variation in Helicobacter pylori lipopolysaccharide due to changes in the lenghts of poly(C) tracts in α3-fucosyltransferase genes. Infect Immun. 1999;67:5361–5366. - PMC - PubMed
-
- Bertram-Drogatz P A, Sobek-Klocke I, Möller C, Wingbermühle D, Beil W, Sewing K-F, Manns M P, Wagner S. Growth characteristics and influence of antibiotics on rough/smooth phenotypic variants of Helicobacter pylori. Eur J Clin Microbiol Infect Dis. 1999;18:490–495. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

