Overexpression, purification and characterization of RecJ protein from Thermus thermophilus HB8 and its core domain
- PMID: 11713311
- PMCID: PMC92510
- DOI: 10.1093/nar/29.22.4617
Overexpression, purification and characterization of RecJ protein from Thermus thermophilus HB8 and its core domain
Abstract
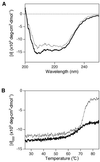
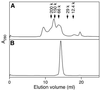
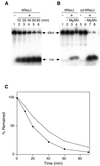
A recJ homolog was cloned from the extremely thermophilic bacterium Thermus themophilus HB8. It encodes a 527 amino acid protein that has 33% identity to Escherichia coli RecJ protein and includes the characteristic motifs conserved among RecJ homologs. Although T.thermophilus RecJ protein (ttRecJ) was expressed as an inclusion body, it was purified in soluble form through denaturation with urea and subsequent refolding steps. Limited proteolysis showed that ttRecJ has a protease-resistant core domain, which includes all the conserved motifs. We constructed a truncated ttRecJ gene that corresponds to the core domain (cd-ttRecJ). cd-ttRecJ was overexpressed in soluble form and purified. ttRecJ and cd-ttRecJ were stable up to 60 degrees C. Size exclusion chromatography indicated that ttRecJ exists in several oligomeric states, whereas cd-ttRecJ is monomeric in solution. Both proteins have 5'-->3' exonuclease activity, which was enhanced by increasing the temperature to 50 degrees C. Mg(2+), Mn(2+) or Co(2+) ions were required to activate both proteins, whereas Ca(2+) and Zn(2+) had no effects.
Figures



Similar articles
-
Thermostable repair enzyme for oxidative DNA damage from extremely thermophilic bacterium, Thermus thermophilus HB8.Nucleic Acids Res. 1998 Feb 15;26(4):903-10. doi: 10.1093/nar/26.4.903. Nucleic Acids Res. 1998. PMID: 9461446 Free PMC article.
-
Mismatch DNA recognition protein from an extremely thermophilic bacterium, Thermus thermophilus HB8.Nucleic Acids Res. 1996 Feb 15;24(4):640-7. doi: 10.1093/nar/24.4.640. Nucleic Acids Res. 1996. PMID: 8604304 Free PMC article.
-
Cloning, sequencing and expression of the uvrA gene from an extremely thermophilic bacterium, Thermus thermophilus HB8.Gene. 1996 May 24;171(1):103-6. doi: 10.1016/0378-1119(96)00052-2. Gene. 1996. PMID: 8675016
-
ATPase activity of UvrB protein form Thermus thermophilus HB8 and its interaction with DNA.J Biol Chem. 1996 Apr 19;271(16):9612-8. doi: 10.1074/jbc.271.16.9612. J Biol Chem. 1996. PMID: 8621636
-
A conserved ancient domain joins the growing superfamily of 3'-5' exonucleases.Curr Biol. 1997 Oct 1;7(10):R604-6. doi: 10.1016/s0960-9822(06)00311-3. Curr Biol. 1997. PMID: 9368736 Review. No abstract available.
Cited by
-
Multiplex Digital MicroRNA Detection Using Cross-Inhibitory DNA Circuits.ACS Sens. 2020 Aug 28;5(8):2430-2437. doi: 10.1021/acssensors.0c00593. Epub 2020 Jul 25. ACS Sens. 2020. PMID: 32602335 Free PMC article.
-
Molecular mechanisms of the whole DNA repair system: a comparison of bacterial and eukaryotic systems.J Nucleic Acids. 2010 Oct 14;2010:179594. doi: 10.4061/2010/179594. J Nucleic Acids. 2010. PMID: 20981145 Free PMC article.
-
Functional analysis of single enzymes combining programmable molecular circuits with droplet-based microfluidics.Nat Nanotechnol. 2024 Jun;19(6):800-809. doi: 10.1038/s41565-024-01617-1. Epub 2024 Feb 26. Nat Nanotechnol. 2024. PMID: 38409552
-
A novel single-stranded DNA-specific 3'-5' exonuclease, Thermus thermophilus exonuclease I, is involved in several DNA repair pathways.Nucleic Acids Res. 2010 Sep;38(17):5692-705. doi: 10.1093/nar/gkq350. Epub 2010 May 10. Nucleic Acids Res. 2010. PMID: 20457749 Free PMC article.
-
Structure of RecJ exonuclease defines its specificity for single-stranded DNA.J Biol Chem. 2010 Mar 26;285(13):9762-9769. doi: 10.1074/jbc.M109.096487. Epub 2010 Feb 2. J Biol Chem. 2010. PMID: 20129927 Free PMC article.
References
-
- Friedberg E.C., Walker,G.C. and Siede,W. (1995) DNA Repair and Mutagenesis. American Society of Microbiology, Washington, DC.
-
- Kowalczykowski S.C. (2000) Initiation of genetic recombination and recombination-dependent replication. Trends Biochem. Sci., 25, 156–165. - PubMed
-
- Cooper D.L., Lahue,R.S. and Modrich,P. (1993) Methyl-directed mismatch repair is bidirectional. J. Biol. Chem., 268, 11823–11829. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

