Analysis of crystal structures of aspartic proteinases: on the role of amino acid residues adjacent to the catalytic site of pepsin-like enzymes
- PMID: 11714911
- PMCID: PMC2374050
- DOI: 10.1110/ps.25801
Analysis of crystal structures of aspartic proteinases: on the role of amino acid residues adjacent to the catalytic site of pepsin-like enzymes
Abstract
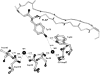
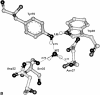
To elucidate the role of amino acid residues adjacent to the catalytic site of pepsin-like enzymes, we analyzed and compared the crystal structures of these enzymes, their complexes with inhibitors, and zymogens in the active site area (a total of 82 structures). In addition to the water molecule (W1) located between the active carboxyls and playing a role of the nucleophile during catalytic reaction, another water molecule (W2) at the vicinity of the active groups was found to be completely conserved. This water molecule plays an essential role in formation of a chain of hydrogen-bonded residues between the active site flap and the active carboxyls on ligand binding. These data suggest a new approach to understanding the role of residues around the catalytic site, which can assist the development of the catalytic reaction. The influence of groups adjacent to the active carboxyls is manifested by pepsin activity at pH 1.0. Some features of pepsin-like enzymes and their mutants are discussed in the framework of the approach.
Figures





Similar articles
-
A new way of looking at aspartic proteinase structures: a comparison of pepsin structure to other aspartic proteinases in the near active site region.Adv Exp Med Biol. 1995;362:19-32. doi: 10.1007/978-1-4615-1871-6_3. Adv Exp Med Biol. 1995. PMID: 8540318 No abstract available.
-
[Conserved interactions of the active carboxyls in pepsin-like enzymes and retroviral proteases].Mol Biol (Mosk). 2002 Sep-Oct;36(5):939-44. Mol Biol (Mosk). 2002. PMID: 12391858 Russian.
-
Revised 2.3 A structure of porcine pepsin: evidence for a flexible subdomain.Proteins. 1990;8(1):62-81. doi: 10.1002/prot.340080109. Proteins. 1990. PMID: 2217165
-
Structural aspects of activation pathways of aspartic protease zymogens and viral 3C protease precursors.Proc Natl Acad Sci U S A. 1999 Sep 28;96(20):10968-75. doi: 10.1073/pnas.96.20.10968. Proc Natl Acad Sci U S A. 1999. PMID: 10500110 Free PMC article. Review.
-
Nepenthesin, a unique member of a novel subfamily of aspartic proteinases: enzymatic and structural characteristics.Curr Protein Pept Sci. 2005 Dec;6(6):513-25. doi: 10.2174/138920305774933259. Curr Protein Pept Sci. 2005. PMID: 16381601 Review.
Cited by
-
Enzymic and structural characterization of nepenthesin, a unique member of a novel subfamily of aspartic proteinases.Biochem J. 2004 Jul 1;381(Pt 1):295-306. doi: 10.1042/BJ20031575. Biochem J. 2004. PMID: 15035659 Free PMC article.
-
Structural studies of vacuolar plasmepsins.Biochim Biophys Acta. 2012 Jan;1824(1):207-23. doi: 10.1016/j.bbapap.2011.04.008. Epub 2011 Apr 20. Biochim Biophys Acta. 2012. PMID: 21540129 Free PMC article. Review.
-
Exploring the binding of BACE-1 inhibitors using comparative binding energy analysis (COMBINE).BMC Struct Biol. 2012 Aug 27;12:21. doi: 10.1186/1472-6807-12-21. BMC Struct Biol. 2012. PMID: 22925713 Free PMC article.
-
Structures of plasmepsin X from Plasmodium falciparum reveal a novel inactivation mechanism of the zymogen and molecular basis for binding of inhibitors in mature enzyme.Protein Sci. 2022 Apr;31(4):882-899. doi: 10.1002/pro.4279. Epub 2022 Feb 5. Protein Sci. 2022. PMID: 35048450 Free PMC article.
-
Exploring the pH-Dependent Structure-Dynamics-Function Relationship of Human Renin.J Chem Inf Model. 2021 Jan 25;61(1):400-407. doi: 10.1021/acs.jcim.0c01201. Epub 2020 Dec 23. J Chem Inf Model. 2021. PMID: 33356221 Free PMC article.
References
-
- Abad-Zapatero, C., Rydel, T.J., and Erickson, J.W. 1990. Revised 2.3 Å structure of porcine pepsin. Evidence for a flexible subdomain. Proteins Struct. Funct. Genet. 8 62–81. - PubMed
-
- Abad-Zapatero, C., Goldman, R., Muchmore, S.W., Hutchins, C., Stewart, K., Navaza, J., Payne, C.D., and Ray, T.E. 1996. Structure of a secreted aspartic proteinase from Candida albicans complexed with a potent inhibitor: Implications for the design of antifungal agents. Protein Sci. 5 640–652. - PMC - PubMed
-
- Aguilar, C.F., Cronin, N.B., Badasso, M., Dreyer, T., Newman, M.P., Cooper, J.B., Hoover, D.J., Wood, S.P., Johnson, M.S., and Blundell, T.L. 1997. The three dimensional structure at 2.4 Å resolution of glycosylated proteinase A from the lysosome-like vacuole of Saccharomyces cerevisiae. J. Mol. Biol. 257 899–915. - PubMed
-
- Andreeva, N.S. and James, M.N.G. 1991. Why does pepsin have a negative charge at very low pH? An analysis of conserved charged residues in aspartic proteinases. Adv. Exp. Med. Biol. 306 39–45. - PubMed
-
- Andreeva, N.S. and Pechik, I.V. 1995. Comparison of three-dimensional structures of flexible protein molecules. Mol. Biol. 22 650–657. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

