Identification of residues critical for metallo-beta-lactamase function by codon randomization and selection
- PMID: 11714924
- PMCID: PMC2374027
- DOI: 10.1110/ps.40884
Identification of residues critical for metallo-beta-lactamase function by codon randomization and selection
Abstract
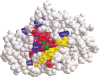
IMP-1 beta-lactamase is a zinc metallo-enzyme encoded by the transferable bla(IMP-1) gene, which confers resistance to virtually all beta-lactam antibiotics including carbapenems. To understand how IMP-1 recognizes and hydrolyzes beta-lactam antibiotics it is important to determine which amino acid residues are critical for catalysis and which residues control substrate specificity. We randomized 27 individual codons in the bla(IMP-1) gene to create libraries that contain all possible amino acid substitutions at residue positions in and near the active site of IMP-1. Mutants from the random libraries were selected for the ability to confer ampicillin resistance to Escherichia coli. Of the positions randomized, >50% do not tolerate amino acid substitutions, suggesting they are essential for IMP-1 function. The remaining positions tolerate amino acid substitutions and may influence the substrate specificity of the enzyme. Interestingly, kinetic studies for one of the functional mutants, Asn233Ala, indicate that an alanine substitution at this position significantly increases catalytic efficiency as compared with the wild-type enzyme.
Figures
Similar articles
-
Analysis of the context dependent sequence requirements of active site residues in the metallo-beta-lactamase IMP-1.J Mol Biol. 2004 Nov 26;344(3):653-63. doi: 10.1016/j.jmb.2004.09.074. J Mol Biol. 2004. PMID: 15533435
-
Insight into stereochemistry of a new IMP allelic variant (IMP-55) metallo-β-lactamase identified in a clinical strain of Acinetobacter baumannii.Infect Genet Evol. 2017 Jul;51:118-126. doi: 10.1016/j.meegid.2017.03.018. Epub 2017 Mar 21. Infect Genet Evol. 2017. PMID: 28336429
-
Structural and biochemical evidence that a TEM-1 beta-lactamase N170G active site mutant acts via substrate-assisted catalysis.J Biol Chem. 2009 Nov 27;284(48):33703-12. doi: 10.1074/jbc.M109.053819. Epub 2009 Oct 6. J Biol Chem. 2009. PMID: 19812041 Free PMC article.
-
Phage display of enzymes and in vitro selection for catalytic activity.Appl Biochem Biotechnol. 1994 May-Jun;47(2-3):175-89; discussion 189-90. doi: 10.1007/BF02787933. Appl Biochem Biotechnol. 1994. PMID: 7944336 Review.
-
Expansion of the zinc metallo-hydrolase family of the beta-lactamase fold.FEBS Lett. 2001 Aug 10;503(1):1-6. doi: 10.1016/s0014-5793(01)02686-2. FEBS Lett. 2001. PMID: 11513844 Review.
Cited by
-
Probing the role of Met221 in the unusual metallo-β-lactamase GOB-18.Inorg Chem. 2012 Nov 19;51(22):12419-25. doi: 10.1021/ic301801h. Epub 2012 Oct 31. Inorg Chem. 2012. PMID: 23113650 Free PMC article.
-
Host-specific enzyme-substrate interactions in SPM-1 metallo-β-lactamase are modulated by second sphere residues.PLoS Pathog. 2014 Jan;10(1):e1003817. doi: 10.1371/journal.ppat.1003817. Epub 2014 Jan 2. PLoS Pathog. 2014. PMID: 24391494 Free PMC article.
-
Fine mapping of the sequence requirements for binding of beta-lactamase inhibitory protein (BLIP) to TEM-1 beta-lactamase using a genetic screen for BLIP function.J Mol Biol. 2009 Jun 5;389(2):401-12. doi: 10.1016/j.jmb.2009.04.028. Epub 2009 Apr 21. J Mol Biol. 2009. PMID: 19389404 Free PMC article.
-
The exchangeability of amino acids in proteins.Genetics. 2005 Aug;170(4):1459-72. doi: 10.1534/genetics.104.039107. Epub 2005 Jun 8. Genetics. 2005. PMID: 15944362 Free PMC article.
-
Evolution of Metallo-β-lactamases: Trends Revealed by Natural Diversity and in vitro Evolution.Antibiotics (Basel). 2014 Jul 1;3(3):285-316. doi: 10.3390/antibiotics3030285. Antibiotics (Basel). 2014. PMID: 25364574 Free PMC article.
References
-
- Amann, E., Brosius, J., and Ptashne, M. 1983. Vectors bearing a hybrid trp–lac promoter useful for regulated expression of cloned genes in Escherichia coli. Gene 25 167–178. - PubMed
-
- Ambler, R.P. 1980. The structure of β-lactamases. Philos. Trans. Roy. Soc. Lond. B Biol. Sci. 289 321–331. - PubMed
-
- Bullock, W.O., Fernandez, J.M., and Short, J.M. 1987. XL1-Blue: A high efficiency plasmid transforming recA Escherichia coli strain with β-galactosidase selection. BioTechniques 5 376–379.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

