Structure of a quinohemoprotein amine dehydrogenase with an uncommon redox cofactor and highly unusual crosslinking
- PMID: 11717396
- PMCID: PMC64671
- DOI: 10.1073/pnas.241429098
Structure of a quinohemoprotein amine dehydrogenase with an uncommon redox cofactor and highly unusual crosslinking
Abstract
The crystal structure of the heterotrimeric quinohemoprotein amine dehydrogenase from Paracoccus denitrificans has been determined at 2.05-A resolution. Within an 82-residue subunit is contained an unusual redox cofactor, cysteine tryptophylquinone (CTQ), consisting of an orthoquinone-modified tryptophan side chain covalently linked to a nearby cysteine side chain. The subunit is surrounded on three sides by a 489-residue, four-domain subunit that includes a diheme cytochrome c. Both subunits sit on the surface of a third subunit, a 337-residue seven-bladed beta-propeller that forms part of the enzyme active site. The small catalytic subunit is internally crosslinked by three highly unusual covalent cysteine to aspartic or glutamic acid thioether linkages in addition to the cofactor crossbridge. The catalytic function of the enzyme as well as the biosynthesis of the unusual catalytic subunit is discussed.
Figures


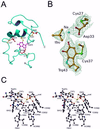


Comment in
-
Building highly sensitive dye assemblies for biosensing from molecular building blocks.Proc Natl Acad Sci U S A. 2001 Dec 18;98(26):14769-72. doi: 10.1073/pnas.251555298. Epub 2001 Dec 11. Proc Natl Acad Sci U S A. 2001. PMID: 11742082 Free PMC article.
-
How many ways to craft a cofactor?Proc Natl Acad Sci U S A. 2001 Dec 18;98(26):14766-8. doi: 10.1073/pnas.011602498. Proc Natl Acad Sci U S A. 2001. PMID: 11752422 Free PMC article. No abstract available.
References
-
- Janes S M, Mu D, Wemmer D, Smith A J, Kaur S, Maltby D, Burlingame A L, Klinman J P. Science. 1990;248:981–987. - PubMed
-
- Wang S X, Mure M, Medzihradszky K F, Burlingame A L, Brown D E, Dooley D M, Smith A J, Kagan H M, Klinman J P. Science. 1996;273:1078–1084. - PubMed
-
- McIntire W S, Wemmer D E, Chistoserdov A, Lidstrom M E. Science. 1991;252:817–824. - PubMed
-
- Klinman J P, Mu D. Annu Rev Biochem. 1994;63:299–344. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

