Evidence that both ligand binding and covalent adaptation drive a two-state equilibrium in the aspartate receptor signaling complex
- PMID: 11723162
- PMCID: PMC2229510
- DOI: 10.1085/jgp.118.6.693
Evidence that both ligand binding and covalent adaptation drive a two-state equilibrium in the aspartate receptor signaling complex
Erratum in
- J Gen Physiol. 2002 Oct;120(4):601.
Abstract
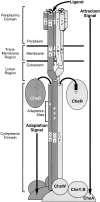
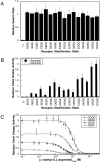
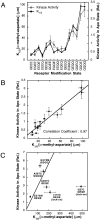
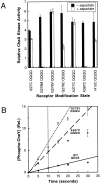
The transmembrane aspartate receptor of bacterial chemotaxis regulates an associated kinase protein in response to both attractant binding to the receptor periplasmic domain and covalent modification of four adaptation sites on the receptor cytoplasmic domain. The existence of at least 16 covalent modification states raises the question of how many stable signaling conformations exist. In the simplest case, the receptor could have just two stable conformations ("on" and "off") yielding the two-state behavior of a toggle-switch. Alternatively, covalent modification could incrementally shift the receptor between many more than two stable conformations, thereby allowing the receptor to function as a rheostatic switch. An important distinction between these models is that the observed functional parameters of a toggle-switch receptor could strongly covary as covalent modification shifts the equilibrium between the on- and off-states, due to population-weighted averaging of the intrinsic on- and off-state parameters. By contrast, covalent modification of a rheostatic receptor would create new conformational states with completely independent parameters. To resolve the toggle-switch and rheostat models, the present study has generated all 16 homogeneous covalent modification states of the receptor adaptation sites, and has compared their effects on the attractant affinity and kinase activity of the reconstituted receptor-kinase signaling complex. This approach reveals that receptor covalent modification modulates both attractant affinity and kinase activity up to 100-fold, respectively. The regulatory effects of individual adaptation sites are not perfectly additive, indicating synergistic interactions between sites. The three adaptation sites at positions 295, 302, and 309 are more important than the site at position 491 in regulating attractant affinity and kinase activity, thereby explaining the previously observed dominance of the former three sites in in vivo studies. The most notable finding is that covalent modification of the adaptation sites alters the receptor attractant affinity and the receptor-regulated kinase activity in a highly correlated fashion, strongly supporting the toggle-switch model. Similarly, certain mutations that drive the receptor into the kinase activating state are found to have correlated effects on attractant affinity. Together these results provide strong evidence that chemotaxis receptors possess just two stable signaling conformations and that the equilibrium between these pure on- and off-states is modulated by both attractant binding and covalent adaptation. It follows that the attractant and adaptation signals drive the same conformational change between the two settings of a toggle. An approach that quantifies the fractional occupancy of the on- and off-states is illustrated.
Figures

Similar articles
-
Quantitative analysis of aspartate receptor signaling complex reveals that the homogeneous two-state model is inadequate: development of a heterogeneous two-state model.J Mol Biol. 2003 Mar 7;326(5):1597-614. doi: 10.1016/s0022-2836(03)00026-3. J Mol Biol. 2003. PMID: 12595268 Free PMC article.
-
Attractant regulation of the aspartate receptor-kinase complex: limited cooperative interactions between receptors and effects of the receptor modification state.Biochemistry. 2000 Aug 8;39(31):9486-93. doi: 10.1021/bi0002737. Biochemistry. 2000. PMID: 10924144 Free PMC article.
-
Engineered socket study of signaling through a four-helix bundle: evidence for a yin-yang mechanism in the kinase control module of the aspartate receptor.Biochemistry. 2009 Oct 6;48(39):9266-77. doi: 10.1021/bi901020d. Biochemistry. 2009. PMID: 19705835 Free PMC article.
-
Adaptation mechanism of the aspartate receptor: electrostatics of the adaptation subdomain play a key role in modulating kinase activity.Biochemistry. 2005 Feb 8;44(5):1550-60. doi: 10.1021/bi048089z. Biochemistry. 2005. PMID: 15683239 Free PMC article.
-
Emerging issues of connexin channels: biophysics fills the gap.Q Rev Biophys. 2001 Aug;34(3):325-472. doi: 10.1017/s0033583501003705. Q Rev Biophys. 2001. PMID: 11838236 Review.
Cited by
-
Excitation and adaptation in bacteria-a model signal transduction system that controls taxis and spatial pattern formation.Int J Mol Sci. 2013 Apr 26;14(5):9205-48. doi: 10.3390/ijms14059205. Int J Mol Sci. 2013. PMID: 23624608 Free PMC article.
-
A "trimer of dimers"-based model for the chemotactic signal transduction network in bacterial chemotaxis.Bull Math Biol. 2012 Oct;74(10):2339-82. doi: 10.1007/s11538-012-9756-7. Epub 2012 Aug 4. Bull Math Biol. 2012. PMID: 22864951 Free PMC article.
-
Receptor-receptor coupling in bacterial chemotaxis: evidence for strongly coupled clusters.Biophys J. 2006 Jun 15;90(12):4317-26. doi: 10.1529/biophysj.105.079905. Epub 2006 Mar 24. Biophys J. 2006. PMID: 16565056 Free PMC article.
-
Quantitative modeling of sensitivity in bacterial chemotaxis: the role of coupling among different chemoreceptor species.Proc Natl Acad Sci U S A. 2003 Jul 8;100(14):8223-8. doi: 10.1073/pnas.1330839100. Epub 2003 Jun 25. Proc Natl Acad Sci U S A. 2003. PMID: 12826616 Free PMC article.
-
Structure, function, and on-off switching of a core unit contact between CheA kinase and CheW adaptor protein in the bacterial chemosensory array: A disulfide mapping and mutagenesis study.Biochemistry. 2013 Nov 5;52(44):7753-65. doi: 10.1021/bi401159k. Epub 2013 Oct 22. Biochemistry. 2013. PMID: 24090207 Free PMC article.
References
-
- Alon U., Surette M.G., Barkai N., Leibler S. Robustness in bacterial chemotaxis. Nature. 1999;397:168–171. - PubMed
-
- Ames P., Parkinson J.S. Transmembrane signaling by bacterial chemoreceptorsE. coli transducers with locked signal output. Cell. 1988;55:817–826. - PubMed
-
- Armitage J.P. Bacterial tactic responses. Adv. Microb. Physiol. 1999;41:229–289. - PubMed
-
- Asakura S., Honda H. Two-state model for bacterial chemoreceptor proteins. The role of multiple methylation. J. Mol. Biol. 1984;176:349–367. - PubMed
-
- Barkai N., Leibler S. Robustness in simple biochemical networks. Nature. 1997;387:913–917. - PubMed

