Candidate odorant receptors from the malaria vector mosquito Anopheles gambiae and evidence of down-regulation in response to blood feeding
- PMID: 11724964
- PMCID: PMC64743
- DOI: 10.1073/pnas.261432998
Candidate odorant receptors from the malaria vector mosquito Anopheles gambiae and evidence of down-regulation in response to blood feeding
Abstract
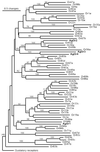
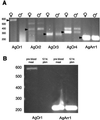
Olfaction plays a major role in host preference and blood feeding, integral behaviors for disease transmission by the malaria vector mosquito Anopheles gambiae sensu stricto (henceforth A. gambiae). We have identified four genes encoding candidate odorant receptors from A. gambiae that are selectively expressed in olfactory organs, contain approximately seven transmembrane domains, and show significant similarity to several putative odorant receptors in Drosophila melanogaster. Furthermore, one of the putative A. gambiae odorant receptors exhibits female-specific antennal expression and is down-regulated 12 h after blood feeding, a period during which substantial reduction in olfactory responses to human odorants has been observed. Taken together, these data suggest these genes encode a family of odorant receptors in A. gambiae, whose further study may aid in the design of novel antimalarial programs.
Figures


References
-
- Hildebrand J G, Shepherd G M. Annu Rev Neurosci. 1997;20:595–631. - PubMed
-
- Takken W. Insect Sci Appl. 1991;12:287–295.
-
- Collins F H, Paskewitz S M. Annu Rev Entomol. 1995;40:195–219. - PubMed
-
- Mombaerts P. Annu Rev Neurosci. 1999;22:487–509. - PubMed
-
- Pilpel Y, Sosinsky A, Lancet D. Essays Biochem. 1998;33:93–104. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

