The genome of swinepox virus
- PMID: 11752168
- PMCID: PMC136851
- DOI: 10.1128/jvi.76.2.783-790.2002
The genome of swinepox virus
Abstract
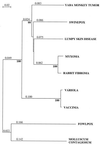
Swinepox virus (SWPV), the sole member of the Suipoxvirus genus of the Poxviridae, is the etiologic agent of a worldwide disease specific for swine. Here we report the genomic sequence of SWPV. The 146-kbp SWPV genome consists of a central coding region bounded by identical 3.7-kbp inverted terminal repeats and contains 150 putative genes. Comparison of SWPV with chordopoxviruses reveals 146 conserved genes encoding proteins involved in basic replicative functions, viral virulence, host range, and immune evasion. Notably, these include genes with similarity to genes for gamma interferon (IFN-gamma) receptor, IFN resistance protein, interleukin-18 binding protein, IFN-alpha/beta binding protein, extracellular enveloped virus host range protein, dUTPase, hydroxysteroid dehydrogenase, superoxide dismutase, serpin, herpesvirus major histocompatibility complex inhibitor, ectromelia virus macrophage host range protein, myxoma virus M011L, variola virus B22R, four ankyrin repeat proteins, three kelch-like proteins, five vaccinia virus (VV) A52R-like family proteins, and two G protein-coupled receptors. The most conserved genomic region is centrally located and corresponds to the VV region located between genes F9L and A38L. Within the terminal 13 kbp, colinearity is disrupted and multiple poxvirus gene homologues are absent or share a lower percentage of amino acid identity. Most of these differences involve genes and gene families with likely functions involving viral virulence and host range. Three open reading frames (SPV018, SPV019. and SPV020) are unique for SWPV. Phylogenetic analysis, genome organization, and amino acid identity indicate that SWPV is most closely related to the capripoxvirus lumpy skin disease virus, followed by the yatapoxvirus yaba-like disease virus and the leporipoxviruses. The gene complement of SWPV better defines Suipoxvirus within the Chordopoxvirinae subfamily and provides a basis for future genetic comparisons.
Figures

References
-
- Antoine, G., F. Scheiflinger, F. Dorner, and F. G. Falkner. 1998. The complete genomic sequence of the modified vaccinia Ankara strain: comparison with other orthopoxviruses. Virology 244:365–396. - PubMed
-
- Barcena, J., M. M. Lorenzo, J. M. Sanchez-Puig, and R. Blasco. 2000. Sequence and analysis of a swinepox virus homologue of the vaccinia virus major envelope protein P37 (F13L). J. Gen. Virol. 81:1073–1085. - PubMed
MeSH terms
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

