HemK, a class of protein methyl transferase with similarity to DNA methyl transferases, methylates polypeptide chain release factors, and hemK knockout induces defects in translational termination
- PMID: 11805295
- PMCID: PMC122215
- DOI: 10.1073/pnas.032488499
HemK, a class of protein methyl transferase with similarity to DNA methyl transferases, methylates polypeptide chain release factors, and hemK knockout induces defects in translational termination
Abstract
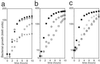
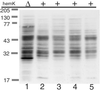
HemK, a universally conserved protein of unknown function, has high amino acid similarity with DNA-(adenine-N6) methyl transferases (MTases). A certain mutation in hemK gene rescues the photosensitive phenotype of a ferrochelatase-deficient (hemH) mutant in Escherichia coli. A hemK knockout strain of E. coli not only suffered severe growth defects, but also showed a global shift in gene expression to anaerobic respiration, as determined by microarray analysis, and this shift may lead to the abrogation of photosensitivity by reducing the oxidative stress. Suppressor mutations that abrogated the growth defects of the hemK knockout strain were isolated and shown to be caused by a threonine to alanine change at codon 246 of polypeptide chain release factor (RF) 2, indicating that hemK plays a role in translational termination. Consistent with such a role, the hemK knockout strain showed an enhanced rate of read-through of nonsense codons and induction of transfer-mRNA-mediated tagging of proteins within the cell. By analysis of the methylation of RF1 and RF2 in vivo and in vitro, we showed that HemK methylates RF1 and RF2 in vitro within the tryptic fragment containing the conserved GGQ motif, and that hemK is required for the methylation within the same fragment of, at least, RF1 in vivo. This is an example of a protein MTase containing the DNA MTase motif and also a protein-(glutamine-N5) MTase.
Figures


Comment in
-
The methylator meets the terminator.Proc Natl Acad Sci U S A. 2002 Feb 5;99(3):1104-6. doi: 10.1073/pnas.042004099. Proc Natl Acad Sci U S A. 2002. PMID: 11830650 Free PMC article. Review. No abstract available.
Similar articles
-
The hemK gene in Escherichia coli encodes the N(5)-glutamine methyltransferase that modifies peptide release factors.EMBO J. 2002 Feb 15;21(4):769-78. doi: 10.1093/emboj/21.4.769. EMBO J. 2002. PMID: 11847124 Free PMC article.
-
The N5-glutamine S-adenosyl-L-methionine-dependent methyltransferase PrmC/HemK in Chlamydia trachomatis methylates class 1 release factors.J Bacteriol. 2005 Jan;187(2):507-11. doi: 10.1128/JB.187.2.507-511.2005. J Bacteriol. 2005. PMID: 15629922 Free PMC article.
-
Cloning and sequencing a HemK-family gene in Porphyromonas gingivalis.DNA Seq. 2003 Feb;14(1):71-4. doi: 10.1080/1042517021000050561. DNA Seq. 2003. PMID: 12751333
-
[Identification and function of protein glutamine (N5) methyl transferase].Tanpakushitsu Kakusan Koso. 2003 May;48(6):733-9. Tanpakushitsu Kakusan Koso. 2003. PMID: 12816008 Review. Japanese. No abstract available.
-
Methylation of class I translation termination factors: structural and functional aspects.Biochimie. 2012 Jul;94(7):1533-43. doi: 10.1016/j.biochi.2012.01.005. Epub 2012 Jan 12. Biochimie. 2012. PMID: 22266024 Review.
Cited by
-
Functional annotation of hypothetical proteins - A review.Bioinformation. 2006 Dec 29;1(8):335-8. doi: 10.6026/97320630001335. Bioinformation. 2006. PMID: 17597916 Free PMC article.
-
Identifying protein function--a call for community action.PLoS Biol. 2004 Mar;2(3):E42. doi: 10.1371/journal.pbio.0020042. Epub 2004 Mar 16. PLoS Biol. 2004. PMID: 15024411 Free PMC article.
-
The Methyltransferase HemK Regulates the Virulence and Nutrient Utilization of the Phytopathogenic Bacterium Xanthomonas citri Subsp. citri.Int J Mol Sci. 2022 Apr 1;23(7):3931. doi: 10.3390/ijms23073931. Int J Mol Sci. 2022. PMID: 35409293 Free PMC article.
-
Molecular Basis of the Slow Growth of Mycoplasma hominis on Different Energy Sources.Front Cell Infect Microbiol. 2022 Jul 7;12:918557. doi: 10.3389/fcimb.2022.918557. eCollection 2022. Front Cell Infect Microbiol. 2022. PMID: 35873139 Free PMC article.
-
Orthogonal Translation for Site-Specific Installation of Post-translational Modifications.Chem Rev. 2024 Mar 13;124(5):2805-2838. doi: 10.1021/acs.chemrev.3c00850. Epub 2024 Feb 19. Chem Rev. 2024. PMID: 38373737 Free PMC article. Review.
References
-
- Nakayashiki T, Nishimura K, Inokuchi H. Gene. 1995;153:67–70. - PubMed
-
- Stover C K, Pham X Q, Erwin A L, Mizoguchi S D, Warrener P, Hickey M J, Brinkman F S, Hufnagle W O, Kowalik D J, Lagrou M, et al. Nature (London) 2000;406:959–964. - PubMed
-
- Presecan E, Moszer I, Boursier L, Cruz Ramos H C, de la Fuente V, Hullo M F, Lelong C, Schleich S, Sekowska A, Song B H, et al. Microbiology. 1997;143:3313–3328. - PubMed
-
- Redenbach M, Kieser H M, Denapaite D, Eichner A, Cullum J, Kinashi H, Hopwood D A. Mol Microbiol. 1996;21:77–96. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

