Formation of transitory intrachain and interchain disulfide bonds accompanies the folding and oligomerization of simian virus 40 Vp1 in the cytoplasm
- PMID: 11805304
- PMCID: PMC122194
- DOI: 10.1073/pnas.032668699
Formation of transitory intrachain and interchain disulfide bonds accompanies the folding and oligomerization of simian virus 40 Vp1 in the cytoplasm
Abstract
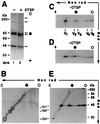
Pentamer formation by Vp1, the major capsid protein of simian virus 40, requires an interdigitation of structural elements from the Vp1 monomers [Liddington, R. C., Yan, Y., Moulai, J., Sahli, R., Benjamin, T. L. & Harrison, S. C. (1991) Nature (London) 354, 278-284]. Our analyses reveal that disulfide-linked Vp1 homooligomers are present in the simian virus 40-infected cytoplasm and that they are derived from a 41-kDa monomeric intermediate containing an intrachain disulfide bond(s). The 41-kDa species, emerging within 5 min of pulse labeling with [(35)S]methionine, is converted into a 45-kDa, disulfide-free Vp1 monomer and disulfide-bonded dimers through pentamers. The covalent oligomer formation is blocked in the presence of a sulfhydryl-modifying reagent. We propose that there are two stages in this Vp1 disulfide bonding. First, the newly synthesized Vp1 monomers acquire intrachain bonds as they fold and begin to interact. Next, these bonds are replaced with intermolecular bonds as the monomers assemble into pentamers. This sequential appearance of transitory disulfide bonds is consistent with a role for sulfhydryl-disulfide redox reactions in the coordinate folding of Vp1 chains into pentamers. The cytoplasmic Vp1 does not colocalize with marker proteins of the endoplasmic reticulum. This paper demonstrates in vivo disulfide formations and exchanges coupled to the folding and oligomerization of a mammalian protein in the cytoplasm, outside the secretory pathway. Such disulfide dynamics may be a general phenomenon for other cysteine-bearing mammalian proteins that fold in the cytoplasm.
Figures

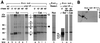

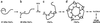
Similar articles
-
Cys9, Cys104 and Cys207 of simian virus 40 Vp1 are essential for inter-pentamer disulfide-linkage and stabilization in cell-free lysates.J Gen Virol. 1999 Sep;80 ( Pt 9):2481-2489. doi: 10.1099/0022-1317-80-9-2481. J Gen Virol. 1999. PMID: 10501505
-
SV40 VP1 assembles into disulfide-linked postpentameric complexes in cell-free lysates.Virology. 1995 Feb 20;207(1):251-4. doi: 10.1006/viro.1995.1073. Virology. 1995. PMID: 7871735
-
Formation of covalently modified folding intermediates of simian virus 40 Vp1 in large T antigen-expressing cells.J Virol. 2013 May;87(9):5053-64. doi: 10.1128/JVI.00955-12. Epub 2013 Feb 20. J Virol. 2013. PMID: 23427157 Free PMC article.
-
Making and breaking disulfide bonds.Annu Rev Microbiol. 1997;51:179-202. doi: 10.1146/annurev.micro.51.1.179. Annu Rev Microbiol. 1997. PMID: 9343348 Review.
-
Simian virus 40 chromatin interaction with the capsid proteins.J Biomol Struct Dyn. 1983 Dec;1(3):689-704. doi: 10.1080/07391102.1983.10507475. J Biomol Struct Dyn. 1983. PMID: 6101085 Review.
Cited by
-
Viral genome structures are optimal for capsid assembly.Elife. 2013 Jun 14;2:e00632. doi: 10.7554/eLife.00632. Elife. 2013. PMID: 23795290 Free PMC article.
-
The C-terminal cysteine annulus participates in auto-chaperone function for Salmonella phage P22 tailspike folding and assembly.Bacteriophage. 2012 Jan 1;2(1):36-49. doi: 10.4161/bact.19775. Bacteriophage. 2012. PMID: 22666655 Free PMC article.
-
Host-membrane interacting interface of the SARS coronavirus envelope protein: Immense functional potential of C-terminal domain.Biophys Chem. 2020 Nov;266:106452. doi: 10.1016/j.bpc.2020.106452. Epub 2020 Aug 11. Biophys Chem. 2020. PMID: 32818817 Free PMC article. Review.
-
Disulfide bonding among micro 1 trimers in mammalian reovirus outer capsid: a late and reversible step in virion morphogenesis.J Virol. 2003 May;77(9):5389-400. doi: 10.1128/jvi.77.9.5389-5400.2003. J Virol. 2003. PMID: 12692241 Free PMC article.
-
Virus strategies for passing the nuclear envelope barrier.Nucleus. 2012 Nov-Dec;3(6):526-39. doi: 10.4161/nucl.21979. Epub 2012 Aug 28. Nucleus. 2012. PMID: 22929056 Free PMC article. Review.
References
-
- Yon J M. Braz J Med Biol Res. 2001;34:419–435. - PubMed
-
- Creighton T E. Biol Chem. 1997;378:731–744. - PubMed
-
- Wedemeyer W J, Welker E, Narayan M, Scheraga H A. Biochemistry. 2000;39:4207–4216. - PubMed
-
- Bergman L W, Kuehl W M. J Biol Chem. 1979;254:5690–5694. - PubMed
-
- Bergman L W, Kuehl W M. J Biol Chem. 1979;254:8869–8876. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

