Intronless homing: site-specific endonuclease SegF of bacteriophage T4 mediates localized marker exclusion analogous to homing endonucleases of group I introns
- PMID: 11825876
- PMCID: PMC155333
- DOI: 10.1101/gad.960302
Intronless homing: site-specific endonuclease SegF of bacteriophage T4 mediates localized marker exclusion analogous to homing endonucleases of group I introns
Abstract
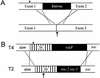
All genetic markers from phage T2 are partially excluded from the progeny of mixed infections with the related phage T4 (general, or phage exclusion). Several loci, including gene 56 of T2, are more dramatically excluded, being present in only approximately 1% of the progeny. This phenomenon is referred to as localized marker exclusion. Gene 69 is adjacent to gene 56 of T4 but is absent in T2, being replaced by completely nonhomologous DNA. We describe SegF, a novel site-specific DNA endonuclease encoded by gene 69, which is similar to GIY-YIG homing endonucleases of group I introns. Interestingly, SegF preferentially cleaves gene 56 of T2, both in vitro and in vivo, compared with that of phage T4. Repair of the double-strand break (DSB) results in the predominance of T4 genes 56 and segF in the progeny, with exclusion of the corresponding T2 sequences. Localized exclusion of T2 gene 56 is dependent on full-length SegF and is likely analogous to group I intron homing, in which repair of a DSB results in coconversion of markers in the flanking DNA. Phage T4 has many optional homing endonuclease genes similar to segF, whereas similar endonuclease genes are relatively rare in other members of the T-even family of bacteriophages. We propose that the general advantage enjoyed by T4 phage, over almost all of its relatives, is a cumulative effect of many of these localized events.
Figures



References
-
- Bell-Pedersen D, Quirk SM, Aubrey M, Belfort M. A site-specific endonuclease and co-conversion of flanking exons associated with the mobile td intron of phage T4. Gene. 1989;82:119–126. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
