Mutation of a chitinase-like gene causes ectopic deposition of lignin, aberrant cell shapes, and overproduction of ethylene
- PMID: 11826306
- PMCID: PMC150558
- DOI: 10.1105/tpc.010278
Mutation of a chitinase-like gene causes ectopic deposition of lignin, aberrant cell shapes, and overproduction of ethylene
Abstract
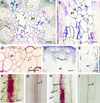
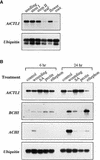
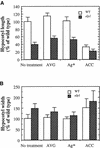
Chitinase-like proteins have long been proposed to play roles in normal plant growth and development, but no mutations in chitinase-like genes have been obtained previously to support this hypothesis. In this study, we have shown that the gene responsible for the elp1 mutation in Arabidopsis encodes a chitinase-like protein (AtCTL1). Mutation of this chitinase-like gene caused ectopic deposition of lignin and aberrant shapes of cells with incomplete cell walls in the pith of inflorescence stems. The AtCTL1 gene was expressed in all organs during normal plant growth and development, but it was not induced by wounding, salicylic acid, pectin fragments, or ethylene. Consistent with its ubiquitous expression pattern, mutation of the AtCTL1 gene affected many aspects of plant growth and development, including exaggerated hook curvature, reduced length and increased diameter of hypocotyls in dark-grown seedlings, and reduced root length and increased number of root hairs in light-grown seedlings. The mutant phenotypes could be rescued partially by ethylene inhibitors, and ethylene production in the mutant was significantly greater than in the wild type. Together, these results suggest that AtCTL1, a chitinase-like gene, is essential for normal plant growth and development in Arabidopsis.
Figures







Similar articles
-
Mutation of the chitinase-like protein-encoding AtCTL2 gene enhances lignin accumulation in dark-grown Arabidopsis seedlings.J Plant Physiol. 2010 May 15;167(8):650-8. doi: 10.1016/j.jplph.2009.12.001. Epub 2010 Jan 6. J Plant Physiol. 2010. PMID: 20056293
-
Arabidopsis hot2 encodes an endochitinase-like protein that is essential for tolerance to heat, salt and drought stresses.Plant J. 2007 Jan;49(2):184-93. doi: 10.1111/j.1365-313X.2006.02950.x. Epub 2006 Dec 6. Plant J. 2007. PMID: 17156413
-
Ectopic deposition of lignin in the pith of stems of two Arabidopsis mutants.Plant Physiol. 2000 May;123(1):59-70. doi: 10.1104/pp.123.1.59. Plant Physiol. 2000. PMID: 10806225 Free PMC article.
-
Chitinase-like protein CTL1 plays a role in altering root system architecture in response to multiple environmental conditions.Plant Physiol. 2010 Feb;152(2):904-17. doi: 10.1104/pp.109.149849. Epub 2009 Dec 9. Plant Physiol. 2010. PMID: 20007445 Free PMC article.
-
Mammalian chitinase-like proteins.EXS. 1999;87:211-21. doi: 10.1007/978-3-0348-8757-1_15. EXS. 1999. PMID: 10906962 Review.
Cited by
-
LysM receptors recognize friend and foe.Proc Natl Acad Sci U S A. 2006 Jul 18;103(29):10829-30. doi: 10.1073/pnas.0604601103. Epub 2006 Jul 10. Proc Natl Acad Sci U S A. 2006. PMID: 16832046 Free PMC article. No abstract available.
-
Accumulation of N-acetylglucosamine oligomers in the plant cell wall affects plant architecture in a dose-dependent and conditional manner.Plant Physiol. 2014 May;165(1):290-308. doi: 10.1104/pp.113.233742. Epub 2014 Mar 24. Plant Physiol. 2014. PMID: 24664205 Free PMC article.
-
Patatin-related phospholipase pPLAIIIβ-induced changes in lipid metabolism alter cellulose content and cell elongation in Arabidopsis.Plant Cell. 2011 Mar;23(3):1107-23. doi: 10.1105/tpc.110.081240. Epub 2011 Mar 29. Plant Cell. 2011. PMID: 21447788 Free PMC article.
-
Three Pectin Methylesterase Inhibitors Protect Cell Wall Integrity for Arabidopsis Immunity to Botrytis.Plant Physiol. 2017 Mar;173(3):1844-1863. doi: 10.1104/pp.16.01185. Epub 2017 Jan 12. Plant Physiol. 2017. PMID: 28082716 Free PMC article.
-
Irritable walls: the plant extracellular matrix and signaling.Plant Physiol. 2010 Jun;153(2):467-78. doi: 10.1104/pp.110.153940. Epub 2010 Feb 12. Plant Physiol. 2010. PMID: 20154095 Free PMC article. Review. No abstract available.
References
-
- Ancillo, G., Witte, B., Schmelzer, E., and Kombrink, E. (1999). A distinct member of the basic (class I) chitinase gene family in potato is specifically expressed in epidermal cells. Plant Mol. Biol. 39, 1137–1151. - PubMed
-
- Bechtold, N., and Bouchez, D. (1994). In planta Agrobacterium-mediated transformation of adult Arabidopsis thaliana plants by vacuum infiltration. In Gene Transfer to Plants, I. Potrykus and G. Spangenberg, eds (Berlin: Springer-Verlag), pp. 19–23.
-
- Benhamou, N., and Asselin, A. (1989). Attempted localization of a substrate for chitinases in plant cells reveals abundant N-acetyl-d-glucosamine residues in secondary walls. Biol. Cell 67, 341–350.
-
- Bent, A.F., Kunkel, B.N., Dahlbeck, D., Brown, K.L., Schmidt, R., Giraudat, J., Leung, J., and Staskawicz, B.J. (1994). RPS2 of Arabidopsis thaliana: A leucine-rich repeat class of plant disease resistance genes. Science 265, 1856–1860. - PubMed
-
- Broglie, K., Chet, I., Hollyday, M., Cressman, R., Biddle, P., Knowlton, S., Mauvais, C.J., and Broglie, R. (1991). Transgenic plants with enhanced resistance to the fungal pathogen Rhizoctonia solani. Science 254, 1194–1197. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

