The tRNA primer activation signal in the human immunodeficiency virus type 1 genome is important for initiation and processive elongation of reverse transcription
- PMID: 11836411
- PMCID: PMC153804
- DOI: 10.1128/jvi.76.5.2329-2339.2002
The tRNA primer activation signal in the human immunodeficiency virus type 1 genome is important for initiation and processive elongation of reverse transcription
Abstract
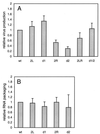
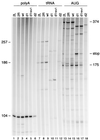
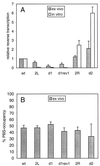
Human immunodeficiency virus type 1 (HIV-1) reverse transcription is primed by the cellular tRNA(3)(Lys) molecule, which binds, with its 3"-terminal 18 nucleotides (nt), to a complementary sequence in the viral genome, the primer-binding site (PBS). Besides PBS-anti-PBS pairing, additional interactions between viral RNA sequences and the tRNA primer are thought to regulate the process of reverse transcription. We previously identified a novel 8-nt sequence motif in the U5 region of the HIV-1 RNA genome that is critical for tRNA(3)(Lys)-mediated initiation of reverse transcription in vitro. This motif activates initiation from the natural tRNA(3)(Lys) primer but is not involved in tRNA placement and was therefore termed primer activation signal (PAS). It was proposed that the PAS interacts with the anti-PAS motif in the TphiC arm of tRNA(3)(Lys). In this study, we analyzed several PAS-mutated viruses and performed reverse transcription assays with virion-extracted RNA-tRNA complexes. Mutation of the PAS reduced the efficiency of tRNA-primed reverse transcription. In contrast, mutations in the opposing leader sequence that trigger release of the PAS from base pairing stimulated reverse transcription. These results are similar to the reverse transcription effects observed in vitro. We also selected revertant viruses that partially overcome the reverse transcription defect of the PAS deletion mutant. Remarkably, all revertants acquired a single nucleotide substitution that does not restore the PAS sequence but that stimulates elongation of reverse transcription. These combined results indicate that the additional PAS-anti-PAS interaction is needed to assemble an initiation-competent and processive reverse transcription complex.
Figures
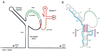





Similar articles
-
Switching the in vitro tRNA usage of HIV-1 by simultaneous adaptation of the PBS and PAS.RNA. 2002 Mar;8(3):357-69. doi: 10.1017/s1355838202028194. RNA. 2002. PMID: 12003495 Free PMC article.
-
A structured RNA motif is involved in correct placement of the tRNA(3)(Lys) primer onto the human immunodeficiency virus genome.J Virol. 2000 Mar;74(5):2227-38. doi: 10.1128/jvi.74.5.2227-2238.2000. J Virol. 2000. PMID: 10666253 Free PMC article.
-
The Interaction between tRNA(Lys) 3 and the primer activation signal deciphered by NMR spectroscopy.PLoS One. 2013 Jun 6;8(6):e64700. doi: 10.1371/journal.pone.0064700. Print 2013. PLoS One. 2013. PMID: 23762248 Free PMC article.
-
tRNA(Lys3): the primer tRNA for reverse transcription in HIV-1.IUBMB Life. 2002 Feb;53(2):107-14. doi: 10.1080/15216540211469. IUBMB Life. 2002. PMID: 12049192 Review.
-
Initiation of HIV-1 reverse transcription and functional role of nucleocapsid-mediated tRNA/viral genome interactions.Virus Res. 2012 Nov;169(2):324-39. doi: 10.1016/j.virusres.2012.06.006. Epub 2012 Jun 18. Virus Res. 2012. PMID: 22721779 Review.
Cited by
-
The three-way junction structure of the HIV-1 PBS-segment binds host enzyme important for viral infectivity.Nucleic Acids Res. 2021 Jun 4;49(10):5925-5942. doi: 10.1093/nar/gkab342. Nucleic Acids Res. 2021. PMID: 33978756 Free PMC article.
-
A Fourth Generation Lentiviral Vector: Simplifying Genomic Gymnastics.Mol Ther. 2017 Aug 2;25(8):1741-1743. doi: 10.1016/j.ymthe.2017.06.005. Mol Ther. 2017. PMID: 28772133 Free PMC article. No abstract available.
-
Human T-cell leukemia virus type 1 uses a specific tRNAPro isodecoder to prime reverse transcription.RNA. 2024 Jul 16;30(8):967-976. doi: 10.1261/rna.080006.124. RNA. 2024. PMID: 38684316 Free PMC article.
-
Mutations in human immunodeficiency virus type 1 nucleocapsid protein zinc fingers cause premature reverse transcription.J Virol. 2008 Oct;82(19):9318-28. doi: 10.1128/JVI.00583-08. Epub 2008 Jul 30. J Virol. 2008. PMID: 18667500 Free PMC article.
-
Structural polymorphism of the HIV-1 leader region explored by computational methods.Nucleic Acids Res. 2005 Dec 20;33(22):7151-63. doi: 10.1093/nar/gki1015. Print 2005. Nucleic Acids Res. 2005. PMID: 16371347 Free PMC article.
References
-
- Arts, E. J., M. Ghosh, P. S. Jacques, B. Ehresmann, and S. F. J. Le Grice. 1996. Restoration of tRNALys,3-primed (−)-strand DNA synthesis to an HIV-1 reverse transcriptase mutant with extended tRNAs. J. Biol. Chem. 271:9054-9061. - PubMed
-
- Arts, E. J., and S. F. J. Le Grice. 1998. Interaction of retroviral reverse transcriptase with template-primer duplexes during replication. Prog. Nucleic Acid Res. Mol. Biol. 58:339-393. - PubMed
-
- Arts, E. J., S. R. Stetor, Y. Li, J. W. Rausch, K. J. Howard, B. Ehresmann, T. W. North, B. M. Wohrl, R. S. Goody, M. A. Wainberg, and S. F. J. Le Grice. 1996. Initiation of (−) strand DNA synthesis from tRNALys3 on lentiviral RNAs: implications of specific HIV-1 RNA-tRNALys3 interactions inhibiting primer utilization by retroviral reverse trancriptases. Proc. Natl. Acad. Sci. USA 93:10063-10068. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

