Drosophila minichromosome maintenance 6 is required for chorion gene amplification and genomic replication
- PMID: 11854416
- PMCID: PMC65653
- DOI: 10.1091/mbc.01-08-0400
Drosophila minichromosome maintenance 6 is required for chorion gene amplification and genomic replication
Abstract
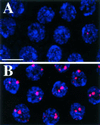
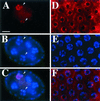
Duplication of the eukaryotic genome initiates from multiple origins of DNA replication whose activity is coordinated with the cell cycle. We have been studying the origins of DNA replication that control amplification of eggshell (chorion) genes during Drosophila oogenesis. Mutation of genes required for amplification results in a thin eggshell phenotype, allowing a genetic dissection of origin regulation. Herein, we show that one mutation corresponds to a subunit of the minichromosome maintenance (MCM) complex of proteins, MCM6. The binding of the MCM complex to origins in G1 as part of a prereplicative complex is critical for the cell cycle regulation of origin licensing. We find that MCM6 associates with other MCM subunits during amplification. These results suggest that chorion origins are bound by an amplification complex that contains MCM proteins and therefore resembles the prereplicative complex. Lethal alleles of MCM6 reveal it is essential for mitotic cycles and endocycles, and suggest that its function is mediated by ATP. We discuss the implications of these findings for the role of MCMs in the coordination of DNA replication during the cell cycle.
Figures






Similar articles
-
The Drosophila chiffon gene is required for chorion gene amplification, and is related to the yeast Dbf4 regulator of DNA replication and cell cycle.Development. 1999 Oct;126(19):4281-93. doi: 10.1242/dev.126.19.4281. Development. 1999. PMID: 10477296
-
Developmental regulation of MCM replication factors in Xenopus laevis.Curr Biol. 1998 Mar 12;8(6):347-50. doi: 10.1016/s0960-9822(98)70136-8. Curr Biol. 1998. PMID: 9512418
-
Chorion gene amplification in Drosophila: A model for metazoan origins of DNA replication and S-phase control.Methods. 1999 Jul;18(3):407-17. doi: 10.1006/meth.1999.0799. Methods. 1999. PMID: 10455001
-
Plant MCM proteins: role in DNA replication and beyond.Plant Mol Biol. 2011 Dec;77(6):537-45. doi: 10.1007/s11103-011-9836-3. Epub 2011 Oct 25. Plant Mol Biol. 2011. PMID: 22038093 Review.
-
Developmental gene amplification and origin regulation.Annu Rev Genet. 2004;38:273-304. doi: 10.1146/annurev.genet.37.110801.143851. Annu Rev Genet. 2004. PMID: 15568978 Review.
Cited by
-
Dm-myb mutant lethality in Drosophila is dependent upon mip130: positive and negative regulation of DNA replication.Genes Dev. 2004 Jul 15;18(14):1667-80. doi: 10.1101/gad.1206604. Genes Dev. 2004. PMID: 15256498 Free PMC article.
-
Genomic markers of ovarian reserve.Semin Reprod Med. 2013 Nov;31(6):399-415. doi: 10.1055/s-0033-1356476. Epub 2013 Oct 7. Semin Reprod Med. 2013. PMID: 24101221 Free PMC article. Review.
-
The origin recognition complex is dispensable for endoreplication in Drosophila.Proc Natl Acad Sci U S A. 2008 Aug 26;105(34):12343-8. doi: 10.1073/pnas.0805189105. Epub 2008 Aug 18. Proc Natl Acad Sci U S A. 2008. PMID: 18711130 Free PMC article.
-
Low levels of p53 protein and chromatin silencing of p53 target genes repress apoptosis in Drosophila endocycling cells.PLoS Genet. 2014 Sep 11;10(9):e1004581. doi: 10.1371/journal.pgen.1004581. eCollection 2014 Sep. PLoS Genet. 2014. PMID: 25211335 Free PMC article.
-
Premature endocycling of Drosophila follicle cells causes pleiotropic defects in oogenesis.Genetics. 2024 Apr 3;226(4):iyae009. doi: 10.1093/genetics/iyae009. Genetics. 2024. PMID: 38302115 Free PMC article.
References
-
- Adams MD, et al. The genome sequence of Drosophila melanogaster. Science. 2000;287:2185–2195. - PubMed
-
- Aparicio O, Weinstein D, Bell S. Components and dynamics of DNA replication complexes in S. cerevisiae: redistribution of MCM proteins and Cdc45p during S phase. Cell. 1997;91:59–69. - PubMed
-
- Bell S, Stillman B. ATP-dependent recognition of eukaryotic origins of DNA replication by a multiprotein complex. Nature. 1992;357:128–134. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

