The evolutionary origin of human subtelomeric homologies--or where the ends begin
- PMID: 11875757
- PMCID: PMC379127
- DOI: 10.1086/339768
The evolutionary origin of human subtelomeric homologies--or where the ends begin
Abstract
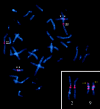
The subtelomeric regions of human chromosomes are comprised of sequence homologies shared between distinct subsets of chromosomes. In the course of developing a set of unique human telomere clones, we identified many clones containing such shared homologies, characterized by the presence of cross-hybridization signals on one or more telomeres in a fluorescence in situ hybridization (FISH) assay. We studied the evolutionary origin of seven subtelomeric clones by performing comparative FISH analysis on a primate panel that included great apes and Old World monkeys. All clones tested showed a single hybridization site in Old World monkeys that corresponded to one of the orthologous human sites, thus indicating the ancestral origin. The timing of the duplication events varied among the subtelomeric regions, from approximately 5 to approximately 25 million years ago. To examine the origin of and mechanism for one of these subtelomeric duplications, we compared the sequence derived from human 2q13--an ancestral fusion site of two great ape telomeric regions--with its paralogous subtelomeric sequences at 9p and 22q. These paralogous regions share large continuous homologies and contain three genes: RABL2B, forkhead box D4, and COBW-like. Our results provide further evidence for subtelomeric-mediated genomic duplication and demonstrate that these segmental duplications are most likely the result of ancestral unbalanced translocations that have been fixed in the genome during recent primate evolution.
Figures
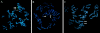



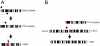

References
Electronic-Database Information
-
- Blast 2 Sequences, http://www.ncbi.nlm.nih.gov/gorf/bl2.html
-
- ClustalW Mutiple Sequence Alignment, http://searchlauncher.bcm.tmc.edu/multi-align/Options/clustalw.html
-
- GenBank, http://www.ncbi.nlm.nih.gov/Genbank/ (for RABL2A [accession number AF095351], RPCI11-395L14 [accession number AL078621], cosmid n94h12 [accession number AC002056], cosmid n1G3 [accession number AC002055], RPCI11-432G15 [accession number AC017074], RPCI11-480C16 [accession number AC016745], RPCI11-65I12 [accession number AC016683], RPCI11-34G16 [accession number AC008268], RPCI11-440D17 [accession number AC009237], RPCI11-403A15 [accession number AL445925], RPCI11-87H9 [accession number AL353763], RPCI11-284A21 [accession number AC025021], RPCI11-88I18 [accession number AL161457], PGM5 [accession number NM_021965], PGMRP [accession number L40933]), FKHL9 [accession number U13223]), COBW-like [accession number AF257330], the physical map of chromosome 22q [accession number NT011526]), and the subtelomeric domain of 4p [accession number Z95704])
-
- Nix Application, http://www.hgmp.mrc.ac.uk/Registered/Webapp/nix/
-
- RepeatMasker Web Server, http://ftp.genome.washington.edu/cgi-bin/RepeatMasker
References
-
- Baldini A, Ried T, Shridhar V, Ogura K, D. Aiuto L, Rocchi M, Ward D (1993) An alphoid DNA sequence conserved in all human and great ape chromosomes: evidence for ancient centromeric sequences at human chromosomal regions 2q21 and 9q13. Hum Genet 90:577–583 - PubMed
-
- Brown WR, MacKinnon PJ, Villasante A, Spurr N, Buckle VJ, Dobson MJ (1990) Structure and polymorphism of human telomere-associated DNA. Cell 63:119–132 - PubMed
Publication types
MeSH terms
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

