Insulin signaling coordinately regulates cardiac size, metabolism, and contractile protein isoform expression
- PMID: 11877471
- PMCID: PMC150890
- DOI: 10.1172/JCI13946
Insulin signaling coordinately regulates cardiac size, metabolism, and contractile protein isoform expression
Abstract
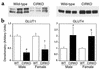
To investigate the role of insulin signaling on postnatal cardiac development, physiology, and cardiac metabolism, we generated mice with a cardiomyocyte-selective insulin receptor knockout (CIRKO) using cre/loxP recombination. Hearts of CIRKO mice were reduced in size by 20-30% due to reduced cardiomyocyte size and had persistent expression of the fetal beta-myosin heavy chain isoform. In CIRKO hearts, glucose transporter 1 (GLUT1) expression was reduced by about 50%, but there was a twofold increase in GLUT4 expression as well as increased rates of cardiac glucose uptake in vivo and increased glycolysis in isolated working hearts. Fatty acid oxidation rates were diminished as a result of reduced expression of enzymes that catalyze mitochondrial beta-oxidation. Although basal rates of glucose oxidation were reduced, insulin unexpectedly stimulated glucose oxidation and glycogenolysis in CIRKO hearts. Cardiac performance in vivo and in isolated hearts was mildly impaired. Thus, insulin signaling plays an important developmental role in regulating postnatal cardiac size, myosin isoform expression, and the switching of cardiac substrate utilization from glucose to fatty acids. Insulin may also modulate cardiac myocyte metabolism through paracrine mechanisms by activating insulin receptors in other cell types within the heart.
Figures






References
-
- Patti ME, Kahn CR. The insulin receptor: a critical link in glucose homeostasis and insulin action. J Basic Clin Physiol Pharmacol. 1998;9:89–109. - PubMed
-
- Bruning JC, et al. Role of brain insulin receptor in control of body weight and reproduction. Science. 2000;289:2122–2125. - PubMed
-
- Drummond-Barbosa D, Spradling AC. Stem cells and their progeny respond to nutritional changes during Drosophila oogenesis. Dev Biol. 2001;231:265–278. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R01 HL043133/HL/NHLBI NIH HHS/United States
- R01 DK092065/DK/NIDDK NIH HHS/United States
- HL-62886/HL/NHLBI NIH HHS/United States
- HL-58073/HL/NHLBI NIH HHS/United States
- P50 HL052338/HL/NHLBI NIH HHS/United States
- DK-43526/DK/NIDDK NIH HHS/United States
- HL-52338/HL/NHLBI NIH HHS/United States
- DK-31036/DK/NIDDK NIH HHS/United States
- DK-33201/DK/NIDDK NIH HHS/United States
- R01 DK043526/DK/NIDDK NIH HHS/United States
- HL-43133/HL/NHLBI NIH HHS/United States
- DK-02495/DK/NIDDK NIH HHS/United States
- R01 DK033201/DK/NIDDK NIH HHS/United States
- R37 DK031036/DK/NIDDK NIH HHS/United States
- R01 HL061483/HL/NHLBI NIH HHS/United States
- R01 DK031036/DK/NIDDK NIH HHS/United States
- HL/AG-61483/HL/NHLBI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

