Structure of a complex of the human alpha/beta T cell receptor (TCR) HA1.7, influenza hemagglutinin peptide, and major histocompatibility complex class II molecule, HLA-DR4 (DRA*0101 and DRB1*0401): insight into TCR cross-restriction and alloreactivity
- PMID: 11877480
- PMCID: PMC2193773
- DOI: 10.1084/jem.20011194
Structure of a complex of the human alpha/beta T cell receptor (TCR) HA1.7, influenza hemagglutinin peptide, and major histocompatibility complex class II molecule, HLA-DR4 (DRA*0101 and DRB1*0401): insight into TCR cross-restriction and alloreactivity
Abstract
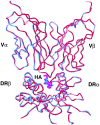
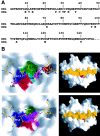
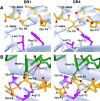
The alpha/beta T cell receptor (TCR) HA1.7 specific for the hemagglutinin (HA) antigen peptide from influenza A virus is HLA-DR1 restricted but cross-reactive for the HA peptide presented by the allo-major histocompatibility complex (MHC) class II molecule HLA-DR4. We report here the structure of the HA1.7/DR4/HA complex, determined by X-ray crystallography at a resolution of 2.4 A. The overall structure of this complex is very similar to the previously reported structure of the HA1.7/DR1/HA complex. Amino acid sequence differences between DR1 and DR4, which are located deep in the peptide binding groove and out of reach for direct contact by the TCR, are able to indirectly influence the antigenicity of the pMHC surface by changing the conformation of HA peptide residues at position P5 and P6. Although TCR HA1.7 is cross-reactive for HA presented by DR1 and DR4 and tolerates these conformational differences, other HA-specific TCRs are sensitive to these changes. We also find a dependence of the width of the MHC class II peptide-binding groove on the sequence of the bound peptide by comparing the HA1.7/DR4/HA complex with the structure of DR4 presenting a collagen peptide. This structural study of TCR cross-reactivity emphasizes how MHC sequence differences can affect TCR binding indirectly by moving peptide atoms.
Figures

Similar articles
-
Minimal conformational plasticity enables TCR cross-reactivity to different MHC class II heterodimers.Sci Rep. 2012;2:629. doi: 10.1038/srep00629. Epub 2012 Sep 4. Sci Rep. 2012. PMID: 22953050 Free PMC article.
-
Structure of a covalently stabilized complex of a human alphabeta T-cell receptor, influenza HA peptide and MHC class II molecule, HLA-DR1.EMBO J. 2000 Nov 1;19(21):5611-24. doi: 10.1093/emboj/19.21.5611. EMBO J. 2000. PMID: 11060013 Free PMC article.
-
Contribution of T-cell receptor-contacting and peptide-binding residues of the class II molecule HLA-DR4 Dw10 to serologic and antigen-specific T-cell recognition.Hum Immunol. 1991 Oct;32(2):110-8. doi: 10.1016/0198-8859(91)90107-k. Hum Immunol. 1991. PMID: 1744001
-
A structural voyage toward an understanding of the MHC-I-restricted immune response: lessons learned and much to be learned.Immunol Rev. 2012 Nov;250(1):61-81. doi: 10.1111/j.1600-065X.2012.01159.x. Immunol Rev. 2012. PMID: 23046123 Review.
-
Structure-function studies of T-cell receptor-superantigen interactions.Immunol Rev. 1998 Jun;163:177-86. doi: 10.1111/j.1600-065x.1998.tb01196.x. Immunol Rev. 1998. PMID: 9700510 Review.
Cited by
-
IMPIPS: the immune protection-inducing protein structure concept in the search for steric-electron and topochemical principles for complete fully-protective chemically synthesised vaccine development.PLoS One. 2015 Apr 16;10(4):e0123249. doi: 10.1371/journal.pone.0123249. eCollection 2015. PLoS One. 2015. PMID: 25879751 Free PMC article.
-
Cytotoxic herpes simplex type 2-specific, DQ0602-restricted CD4 T+-cell clones show alloreactivity to DQ0601.Immunology. 2006 Mar;117(3):350-7. doi: 10.1111/j.1365-2567.2005.02308.x. Immunology. 2006. PMID: 16476054 Free PMC article.
-
Identification of an altered peptide ligand based on the endogenously presented, rheumatoid arthritis-associated, human cartilage glycoprotein-39(263-275) epitope: an MHC anchor variant peptide for immune modulation.Arthritis Res Ther. 2007;9(4):R71. doi: 10.1186/ar2269. Arthritis Res Ther. 2007. PMID: 17645792 Free PMC article.
-
A high-throughput yeast display approach to profile pathogen proteomes for MHC-II binding.Elife. 2022 Jul 4;11:e78589. doi: 10.7554/eLife.78589. Elife. 2022. PMID: 35781135 Free PMC article.
-
Predicting peptide binding affinities to MHC molecules using a modified semi-empirical scoring function.PLoS One. 2011;6(9):e25055. doi: 10.1371/journal.pone.0025055. Epub 2011 Sep 22. PLoS One. 2011. PMID: 21966412 Free PMC article.
References
-
- Ding, Y.H., K.J. Smith, D.N. Garboczi, U. Utz, W.E. Biddison, and D.C. Wiley. 1998. Two human T cell receptors bind in a similar diagonal mode to the HLA-A2/Tax peptide complex using different TCR amino acids. Immunity. 8:403–411. - PubMed
-
- Garboczi, D.N., P. Ghosh, U. Utz, Q.R. Fan, W.E. Biddison, and D.C. Wiley. 1996. Structure of the complex between human T-cell receptor, viral peptide and HLA-A2. Nature. 384:134–141. - PubMed
-
- Garcia, K.C., M. Degano, L.R. Pease, M. Huang, P.A. Peterson, L. Teyton, and I.A. Wilson. 1998. Structural basis of plasticity in T cell receptor recognition of a self peptide-MHC antigen. Science. 279:1166–1172. - PubMed
-
- Reinherz, E.L., K. Tan, L. Tang, P. Kern, J. Liu, Y. Xiong, R.E. Hussey, A. Smolyar, B. Hare, R. Zhang, et al. 1999. The crystal structure of a T cell receptor in complex with peptide and MHC class II. Science. 286:1913–1921. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

