Human immunodeficiency virus type 1 group m protease in cameroon: genetic diversity and protease inhibitor mutational features
- PMID: 11880402
- PMCID: PMC120267
- DOI: 10.1128/JCM.40.3.837-845.2002
Human immunodeficiency virus type 1 group m protease in cameroon: genetic diversity and protease inhibitor mutational features
Abstract
To establish a baseline for monitoring resistance to protease inhibitors (PIs) and examining the efficacy of their use among persons in Cameroon infected with human immunodeficiency virus type 1 (HIV-1), we analyzed genetic variability and PI resistance-associated substitutions in PCR-amplified protease (PR) sequences in strains isolated from 110 HIV-1-infected, drug-naïve Cameroonians. Of the 110 strains, 85 were classified into six HIV-1 PR subtypes, A (n = 1), B (n = 1), F (n = 4), G (n = 7), H (n = 1), and J (n = 7), and a circulating recombinant form, CRF02-AG (n = 64). PR genes from the remaining 25 (23%) specimens were unclassifiable, whereas 2% (7 of 301) unclassifiable PR sequences were reported for a global collection. Two major PI resistance-associated mutations, 20M and 24I, were detected in strains from only two specimens, whereas secondary mutations were found in strains from all samples except one strain of subtype B and two strains of CRF02-AG. The secondary mutations showed the typical PI resistance-associated pattern for non-subtype B viruses in both classifiable and unclassifiable PR genes, with 36I being the predominant (99%) mutation, followed by 63P (18%), 20R (15%), 77I (13%), and 10I or 10V (11%). Of these mutations, dual and triple PI resistance-associated substitutions were found in 38% of all the Cameroonian strains. Compared with classifiable PR sequences, unclassifiable sequences had significantly more dual and triple substitutions (64% versus 30%; P = 0.004). Phenotypic and clinical evaluations are needed to estimate whether PI resistance during antiretroviral drug treatment occurs more rapidly in individuals infected with HIV-1 strains harboring multiple PI resistance-associated substitutions. This information may be important for determination of appropriate drug therapies for HIV-1-infected persons in Cameroon, where more than one-third of HIV-1 strains were found to carry dual and triple minor PI resistance-associated mutations.
Figures


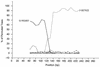


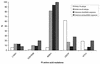
References
-
- Adje, C., R. Cheingsong, T. H. Roels, C. Maurice, G. Djomand, W. Verbiest, K. Hertogs, B. Larder, B. Monga, M. Peeters, S. Eholie, E. Bissagene, M. Coulibaly, R. Respess, S. Z. Wiktor, T. Chorba, J. N. Nkengasong, and The UNAIDS HIV Drug Access Initiative, Abidjan, Cote d'Ivoire. 2001. High prevalence of genotypic and phenotypic HIV-1 drug-resistant strains among patients receiving antiretroviral therapy in Abidjan, Cote d'Ivoire. J. Acquir. Immune Defic. Syndr. 26:501-506. - PubMed
-
- Birk, M., and A. Sonnerborg. 1998. Variations in HIV-1 pol gene associated with reduced sensitivity to antiretroviral drugs in treatment-naïve patients. AIDS 12:2369-2375. - PubMed
-
- Descamps, D., G. Collin, F. Letourneur, C. Apetrei, F. Damond, I. Loussert-Ajaka, F. Simon, S. Saragosti, and F. Brun-Vezinet. 1997. Susceptibility of human immunodeficiency virus type 1 group O isolates to antiretroviral agents: in vitro phenotypic and genotypic analyses. Susceptibility of human immunodeficiency virus type 1 group O isolates to antiretroviral agents: in vitro phenotypic and genotypic analyses. J. Virol. 71:8893-8898. - PMC - PubMed
-
- Descamps, D., C. Apetrei, G. Collin, F. Damond, F. Simon, and F. Brun-Vezinet. 1998. Naturally occurring decreased susceptibility of HIV-1 subtype G to protease inhibitors. AIDS 12:1109-1111. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

