E1 protein of bovine papillomavirus type 1 interferes with E2 protein-mediated tethering of the viral DNA to mitotic chromosomes
- PMID: 11884568
- PMCID: PMC136052
- DOI: 10.1128/jvi.76.7.3440-3451.2002
E1 protein of bovine papillomavirus type 1 interferes with E2 protein-mediated tethering of the viral DNA to mitotic chromosomes
Abstract
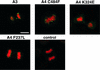
Eukaryotic viruses can maintain latency in dividing cells as extrachromosomal plasmids. It is therefore of vital importance for viruses to ensure nuclear retention and proper segregation of their viral DNA. The bovine papillomavirus (BPV) E2 enhancer protein plays a key role in these processes by tethering the viral DNA to the host cell chromosomes. Viral genomes that harbor phosphorylation mutations in the E2 gene are transformation defective, and for these mutant genomes, neither the viral DNA nor the E2 protein is detected on mitotic chromosomes, while other key functions of E2 in transcription and replication were wild type. Moreover, secondary mutations in both the E2 and E1 proteins lead to suppression of the phosphorylation mutant phenotype and resulted in reattachment of the viral DNA and the E2 protein onto mitotic chromosomes, suggesting that E1 also plays a role in viral genome partitioning. The E1 protein was cytologically always excluded from mitotic chromatin, either as a suppressor allele or as the wild type. In the absence of other viral proteins, an E2 protein containing alanine substitutions for phosphorylation substrates in the hinge region (E2-A4) was detected as wild-type on mitotic chromosomes. However, when wild-type E1 protein levels were increased in cells expressing either the A4 mutant E2 proteins or wild-type E2, the E2-A4 protein was much more sensitive to chromosomal dislocation than was the wild-type protein. In contrast, suppressor alleles of E1 were not capable of such abrogation of E2 binding (A4 or wild-type) to chromosomes. These results suggest that wild-type E1 can be a negative regulator of the chromosomal attachment of E2.
Figures








References
-
- Allikas, A., D. Ord, R. Kurg, S. Kivi, and M. Ustav. 2001. Roles of the hinge region and the DNA binding domain of the bovine papillomavirus type 1 E2 protein in initiation of DNA replication. Virus Res. 75:95-106. - PubMed
-
- Bastien, N., and A. A. McBride. 2000. Interaction of the papillomavirus E2 protein with mitotic chromosomes. Virology 270:124-134. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

