AXR1-ECR1-dependent conjugation of RUB1 to the Arabidopsis Cullin AtCUL1 is required for auxin response
- PMID: 11884684
- PMCID: PMC152922
- DOI: 10.1105/tpc.010282
AXR1-ECR1-dependent conjugation of RUB1 to the Arabidopsis Cullin AtCUL1 is required for auxin response
Abstract
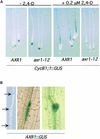
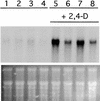
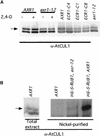
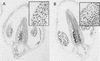
Mutations in the AXR1 gene result in a reduction in auxin response and diverse defects in auxin-regulated growth and development. In a previous study, we showed that AXR1 forms a heterodimer with the ECR1 protein. This enzyme activates the ubiquitin-related protein RUB1 in vitro. Furthermore, we showed that the Skp1-Cul1/Cdc53-F-box (SCF) subunit AtCUL1 is modified by RUB1 in vivo. In this report, we demonstrate that the formation of RUB-AtCUL1 is dependent on AXR1 and ECR1 in vivo. The expression of AXR1 and ECR1 is restricted to zones of active cell division and cell elongation, consistent with their role in growth regulation. These results provide strong support for a model in which RUB conjugation of AtCUL1 affects the function of SCF E3s that are required for auxin response.
Figures



References
-
- Abel, S., Nguyen, M.D., and Theologis, A. (1995). The PS-IAA4/ 5-like family of early auxin-inducible mRNAs in Arabidopsis thaliana. J. Mol. Biol. 251, 533–549. - PubMed
-
- Ausubel, F.M., Brent, R., Kingston, R.E., Moore, D.D., Seidman, J.G., Smith, J.A., and Struhl, K. (1990). Current Protocols in Molecular Biology. (New York: Wiley).
-
- Bechtold, N., Ellis, J., and Pelletier, G. (1993). In planta Agrobacterium mediated gene transfer by infiltration of adult Arabidopsis thaliana plants. C. R. Acad. Sci. Paris Life Sci. 316, 15–18.
-
- Berleth, T., Mattsson, J., and Hardtke, C.S. (2000). Vascular continuity and auxin signals. Trends Plant Sci. 5, 387–393. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

