Structure-based analysis of RNA polymerase function: the largest subunit's rudder contributes critically to elongation complex stability and is not involved in the maintenance of RNA-DNA hybrid length
- PMID: 11889042
- PMCID: PMC125355
- DOI: 10.1093/emboj/21.6.1369
Structure-based analysis of RNA polymerase function: the largest subunit's rudder contributes critically to elongation complex stability and is not involved in the maintenance of RNA-DNA hybrid length
Abstract
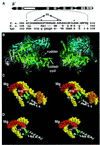
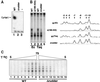
Analysis of multisubunit RNA polymerase (RNAP) structures revealed several elements that may constitute the enzyme's functional sites. One such element, the 'rudder', is formed by an evolutionarily conserved segment of the largest subunit of RNAP and contacts the nascent RNA at the upstream edge of the RNA-DNA hybrid, where the DNA template strand separates from the RNA transcript and re-anneals with the non-template strand. Thus, the rudder could (i) maintain the correct length of the RNA-DNA hybrid; (ii) stabilize the nascent RNA in the complex; and (iii) promote or maintain localized DNA melting at the upstream edge of the bubble. We generated a recombinant RNAP mutant that lacked the rudder and studied its properties in vitro. Our results demonstrate that the rudder is not required for establishment of the upstream boundary of the transcription bubble during promoter complex formation, nor is it required for separation of the nascent RNA from the DNA template strand or transcription termination. Our results suggest that the rudder makes critical contributions to elongation complex stability through direct interactions with the nascent RNA.
Figures


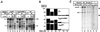

References
-
- Allison L.A., Moyle,M., Shales,M. and Ingles,C.J. (1985) Extensive homology among the largest subunits of eukaryotic and prokaryotic RNA polymerases. Cell, 42, 599–610. - PubMed
-
- Borukhov S. and Goldfarb,A. (1993) Recombinant Escherichia coli RNA polymerase: purification of individually overexpressed subunits and in vitro assembly. Protein Expr. Purif., 4, 503–511. - PubMed
-
- Burgess R.R., Arthur,T.M. and Pietz,B.C. (1998) Interaction of Escherichia coli σ70 with core RNA polymerase. Cold Spring Harb. Symp. Quant. Biol., 63, 277–287. - PubMed
-
- Chamberlin M.J. (1992) New models for the mechanism of transcription elongation and its regulation. Harvey Lect., 88, 1–21. - PubMed
-
- Cramer P., Bushnell,D.A. and Kornberg,R.D. (2001) Structural basis of transcription: RNA polymerase II at 2.8 Å resolution. Science, 292, 1863–1876. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

