Protein unfolding: rigidity lost
- PMID: 11891336
- PMCID: PMC122559
- DOI: 10.1073/pnas.062492699
Protein unfolding: rigidity lost
Abstract
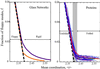
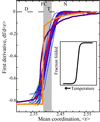
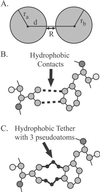
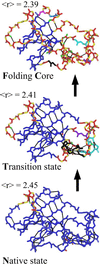
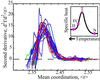
We relate the unfolding of a protein to its loss of structural stability or rigidity. Rigidity and flexibility are well defined concepts in mathematics and physics, with a body of theorems and algorithms that have been applied successfully to materials, allowing the constraints in a network to be related to its deformability. Here we simulate the weakening or dilution of the noncovalent bonds during protein unfolding, and identify the emergence of flexible regions as unfolding proceeds. The transition state is determined from the inflection point in the change in the number of independent bond-rotational degrees of freedom (floppy modes) of the protein as its mean atomic coordination decreases. The first derivative of the fraction of floppy modes as a function of mean coordination is similar to the fraction-folded curve for a protein as a function of denaturant concentration or temperature. The second derivative, a specific heat-like quantity, shows a peak around a mean coordination of <r> = 2.41 for the 26 diverse proteins we have studied. As the protein denatures, it loses rigidity at the transition state, proceeds to a state where just the initial folding core remains stable, then becomes entirely denatured or flexible. This universal behavior for proteins of diverse architecture, including monomers and oligomers, is analogous to the rigid to floppy phase transition in network glasses. This approach provides a unifying view of the phase transitions of proteins and glasses, and identifies the mean coordination as the relevant structural variable, or reaction coordinate, along the unfolding pathway.
Figures
Similar articles
-
Identifying protein folding cores from the evolution of flexible regions during unfolding.J Mol Graph Model. 2002 Dec;21(3):195-207. doi: 10.1016/s1093-3263(02)00146-8. J Mol Graph Model. 2002. PMID: 12463638
-
Protein flexibility predictions using graph theory.Proteins. 2001 Aug 1;44(2):150-65. doi: 10.1002/prot.1081. Proteins. 2001. PMID: 11391777
-
Constraint Network Analysis (CNA): a Python software package for efficiently linking biomacromolecular structure, flexibility, (thermo-)stability, and function.J Chem Inf Model. 2013 Apr 22;53(4):1007-15. doi: 10.1021/ci400044m. Epub 2013 Apr 8. J Chem Inf Model. 2013. PMID: 23517329
-
Engineering more stable proteins.Chem Soc Rev. 2018 Dec 21;47(24):9026-9045. doi: 10.1039/c8cs00014j. Epub 2018 Oct 11. Chem Soc Rev. 2018. PMID: 30306986 Review.
-
Understanding the folding rates and folding nuclei of globular proteins.Curr Protein Pept Sci. 2007 Dec;8(6):521-36. doi: 10.2174/138920307783018695. Curr Protein Pept Sci. 2007. PMID: 18220841 Review.
Cited by
-
Elucidating quantitative stability/flexibility relationships within thioredoxin and its fragments using a distance constraint model.J Mol Biol. 2006 May 5;358(3):882-904. doi: 10.1016/j.jmb.2006.02.015. Epub 2006 Feb 24. J Mol Biol. 2006. PMID: 16542678 Free PMC article.
-
Large scale rigidity-based flexibility analysis of biomolecules.Struct Dyn. 2016 Feb 18;3(1):012005. doi: 10.1063/1.4942414. eCollection 2016 Jan. Struct Dyn. 2016. PMID: 26958583 Free PMC article.
-
Ligand-mediated and tertiary interactions cooperatively stabilize the P1 region in the guanine-sensing riboswitch.PLoS One. 2017 Jun 22;12(6):e0179271. doi: 10.1371/journal.pone.0179271. eCollection 2017. PLoS One. 2017. PMID: 28640851 Free PMC article.
-
Nullspace Sampling with Holonomic Constraints Reveals Molecular Mechanisms of Protein Gαs.PLoS Comput Biol. 2015 Jul 28;11(7):e1004361. doi: 10.1371/journal.pcbi.1004361. eCollection 2015 Jul. PLoS Comput Biol. 2015. PMID: 26218073 Free PMC article.
-
Biochemical and Structural Characterization of Enolase from Chloroflexus aurantiacus: Evidence for a Thermophilic Origin.Front Bioeng Biotechnol. 2015 Jun 1;3:74. doi: 10.3389/fbioe.2015.00074. eCollection 2015. Front Bioeng Biotechnol. 2015. PMID: 26082925 Free PMC article.
References
-
- Bryngelson J D, Onuchic J N, Socci N D, Wolynes P G. Proteins Struct Funct Genet. 1995;21:167–195. - PubMed
-
- Honig B. J Mol Biol. 1999;293:283–293. - PubMed
-
- Radford S E. Trends Biochem Sci. 2000;25:611–618. - PubMed
-
- Baker D. Nature (London) 2000;405:39–42. - PubMed
-
- Jackson S E. Folding Des. 1998;3:R81–R91. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

