HIV-1 integrase interaction with U3 and U5 terminal sequences in vitro defined using substrates with random sequences
- PMID: 11897790
- PMCID: PMC2769074
- DOI: 10.1074/jbc.M201354200
HIV-1 integrase interaction with U3 and U5 terminal sequences in vitro defined using substrates with random sequences
Abstract
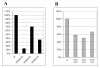
Successful integration of viral genome into a host chromosome depends on interaction between viral integrase and its recognition sequences. We have used a reconstituted concerted human immunodeficiency virus, type 1 (HIV-1), integration system to analyze the role of integrase (IN) recognition sequences in formation of the IN-viral DNA complex capable of concerted integration. HIV-1 integrase was presented with substrates that contained all 4 bases at 8 mismatched positions that define the inverted repeat relationship between U3 and U5 long terminal repeats (LTR) termini and at positions 17-19, which are conserved in the termini. Evidence presented indicates that positions 17-20 of the IN recognition sequences are needed for a concerted DNA integration mechanism. All 4 bases were found at each randomized position in sequenced concerted DNA integrants, although in some instances there were preferences for specific bases. These results indicate that integrase tolerates a significant amount of plasticity as to what constitutes an IN recognition sequence. By having several positions randomized, the concerted integrants were examined for statistically significant relationships between selections of bases at different positions. The results of this analysis show not only relationships between different positions within the same LTR end but also between different positions belonging to opposite DNA termini.
Figures
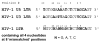

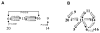

References
-
- Farnet CM, Bushman FD. Cell. 1997;88:483–492. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

