The Dun1 checkpoint kinase phosphorylates and regulates the ribonucleotide reductase inhibitor Sml1
- PMID: 11904430
- PMCID: PMC122595
- DOI: 10.1073/pnas.062502299
The Dun1 checkpoint kinase phosphorylates and regulates the ribonucleotide reductase inhibitor Sml1
Abstract
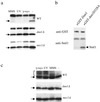
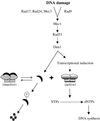
Cell cycle checkpoints are evolutionarily conserved surveillance systems that protect genomic stability and prevent oncogenesis in mammals. One important target of checkpoint control is ribonucleotide reductase (RNR), which catalyzes the rate-limiting step in dNTP and DNA synthesis. In both yeast and humans, RNR is transcriptionally induced after DNA damage via Mec1/Rad53 (yeast) and ATM/CHK2 (human) checkpoint pathways. In addition, yeast checkpoint proteins Mec1 and Rad53 also regulate the RNR inhibitor Sml1. After DNA damage or at S phase, Mec1 and Rad53 control the phosphorylation and concomitant degradation of Sml1 protein. This new layer of control contributes to the increased dNTP production likely necessary for DNA repair and replication; however, the molecular mechanism is unclear. Here we show that Dun1, a downstream kinase of Mec1/Rad53, genetically and physically interacts with Sml1 in vivo. The absence of Dun1 activity leads to the accumulation of Sml1 protein at S phase and after DNA damage. As a result, dun1Delta strains need more time to finish DNA replication, are defective in mitochondrial DNA propagation, and are sensitive to DNA-damaging agents. Moreover, phospho-Sml1 is absent or dramatically reduced in dun1Delta cells. Finally, Dun1 can phosphorylate Sml1 in vitro. These results suggest that Dun1 kinase function is the last step required in the Mec1/Rad53 cascade to remove Sml1 during S phase and after DNA damage.
Figures


Similar articles
-
The ribonucleotide reductase inhibitor Sml1 is a new target of the Mec1/Rad53 kinase cascade during growth and in response to DNA damage.EMBO J. 2001 Jul 2;20(13):3544-53. doi: 10.1093/emboj/20.13.3544. EMBO J. 2001. PMID: 11432841 Free PMC article.
-
Yeast Dun1 kinase regulates ribonucleotide reductase inhibitor Sml1 in response to iron deficiency.Mol Cell Biol. 2014 Sep;34(17):3259-71. doi: 10.1128/MCB.00472-14. Epub 2014 Jun 23. Mol Cell Biol. 2014. PMID: 24958100 Free PMC article.
-
Mutational and structural analyses of the ribonucleotide reductase inhibitor Sml1 define its Rnr1 interaction domain whose inactivation allows suppression of mec1 and rad53 lethality.Mol Cell Biol. 2000 Dec;20(23):9076-83. doi: 10.1128/MCB.20.23.9076-9083.2000. Mol Cell Biol. 2000. PMID: 11074005 Free PMC article.
-
The S-phase checkpoint: targeting the replication fork.Biol Cell. 2009 Aug 19;101(11):617-27. doi: 10.1042/BC20090053. Biol Cell. 2009. PMID: 19686094 Review.
-
Phosphoregulation of the checkpoint kinase Mec1ATR.DNA Repair (Amst). 2023 Sep;129:103543. doi: 10.1016/j.dnarep.2023.103543. Epub 2023 Jul 17. DNA Repair (Amst). 2023. PMID: 37480741 Review.
Cited by
-
Prevention of DNA Rereplication Through a Meiotic Recombination Checkpoint Response.G3 (Bethesda). 2016 Dec 7;6(12):3869-3881. doi: 10.1534/g3.116.033910. G3 (Bethesda). 2016. PMID: 27678521 Free PMC article.
-
The Slx5-Slx8 complex affects sumoylation of DNA repair proteins and negatively regulates recombination.Mol Cell Biol. 2007 Sep;27(17):6153-62. doi: 10.1128/MCB.00787-07. Epub 2007 Jun 25. Mol Cell Biol. 2007. PMID: 17591698 Free PMC article.
-
Analyzing the dose-dependence of the Saccharomyces cerevisiae global transcriptional response to methyl methanesulfonate and ionizing radiation.BMC Genomics. 2006 Dec 1;7:305. doi: 10.1186/1471-2164-7-305. BMC Genomics. 2006. PMID: 17140446 Free PMC article.
-
Inhibition of spindle extension through the yeast S phase checkpoint is coupled to replication fork stability and the integrity of centromeric DNA.Mol Biol Cell. 2019 Oct 15;30(22):2771-2789. doi: 10.1091/mbc.E19-03-0156. Epub 2019 Sep 11. Mol Biol Cell. 2019. PMID: 31509480 Free PMC article.
-
Sml1 Inhibits the DNA Repair Activity of Rev1 in Saccharomyces cerevisiae during Oxidative Stress.Appl Environ Microbiol. 2020 Mar 18;86(7):e02838-19. doi: 10.1128/AEM.02838-19. Print 2020 Mar 18. Appl Environ Microbiol. 2020. PMID: 32005731 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

