The common retroviral insertion locus Dsi1 maps 30 kilobases upstream of the P1 promoter of the murine Runx3/Cbfa3/Aml2 gene
- PMID: 11932403
- PMCID: PMC155108
- DOI: 10.1128/jvi.76.9.4364-4369.2002
The common retroviral insertion locus Dsi1 maps 30 kilobases upstream of the P1 promoter of the murine Runx3/Cbfa3/Aml2 gene
Abstract
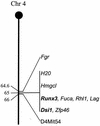
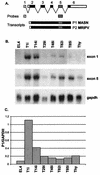
The Dsi1 locus was identified as a common integration site for Moloney murine leukemia virus (MLV) in rat thymic lymphomas, but previous efforts to identify a gene affected by these insertions were unsuccessful. We considered the Runx3 gene a potential candidate on the basis of genetic mapping which showed that Dsi1 and Runx3 are closely linked on mouse chromosome 4 and the precedent of the related Runx2 gene, which emerged recently as a Myc-collaborating gene activated by retroviral insertion in thymic lymphomas of CD2-MYC mice. We now report the physical mapping of the Dsi1 locus to a site 30 kb upstream of the distal (P1) promoter of the murine Runx3 gene. Comparison with the syntenic region of human chromosome 1 shows that the next gene is over 250 kb 5' to Runx3, suggesting that Runx3 may be the primary target of retroviral insertions at Dsi1. Screening of CD2-MYC lymphomas for rearrangements at Dsi1 revealed a tumor cell line harboring an MLV provirus at this locus, in the orientation opposite that of Runx3. Proviral insertion was associated with very high levels of expression of Runx3, with a preponderance of transcripts arising at the P1 promoter. These results confirm that Runx3 is a target of retroviral insertions at Dsi1 and indicate that Runx3 can act as an alternative to Runx2 as a Myc-collaborating gene in thymic lymphoma.
Figures

Similar articles
-
Runx3 knockouts and stomach cancer.EMBO Rep. 2003 Jun;4(6):560-4. doi: 10.1038/sj.embor.embor868. EMBO Rep. 2003. PMID: 12776174 Free PMC article. Review.
-
til-1: a novel proviral insertion locus for Moloney murine leukaemia virus in lymphomas of CD2-myc transgenic mice.J Gen Virol. 1996 Mar;77 ( Pt 3):443-6. doi: 10.1099/0022-1317-77-3-443. J Gen Virol. 1996. PMID: 8601779
-
Activation of multiple genes by provirus integration in the Mlvi-4 locus in T-cell lymphomas induced by Moloney murine leukemia virus.J Virol. 1990 May;64(5):2236-44. doi: 10.1128/JVI.64.5.2236-2244.1990. J Virol. 1990. PMID: 1691313 Free PMC article.
-
Runx2: a novel oncogenic effector revealed by in vivo complementation and retroviral tagging.Oncogene. 2001 Jan 18;20(3):295-302. doi: 10.1038/sj.onc.1204090. Oncogene. 2001. PMID: 11313958
-
The Runx genes as dominant oncogenes.Blood Cells Mol Dis. 2003 Mar-Apr;30(2):194-200. doi: 10.1016/s1079-9796(03)00031-7. Blood Cells Mol Dis. 2003. PMID: 12732183 Review.
Cited by
-
Runx2 disruption promotes immortalization and confers resistance to oncogene-induced senescence in primary murine fibroblasts.Cancer Res. 2007 Dec 1;67(23):11263-71. doi: 10.1158/0008-5472.CAN-07-3016. Cancer Res. 2007. PMID: 18056452 Free PMC article.
-
Addiction to Runx1 is partially attenuated by loss of p53 in the Eµ-Myc lymphoma model.Oncotarget. 2016 Apr 26;7(17):22973-87. doi: 10.18632/oncotarget.8554. Oncotarget. 2016. PMID: 27056890 Free PMC article.
-
Runx1 Orchestrates Sphingolipid Metabolism and Glucocorticoid Resistance in Lymphomagenesis.J Cell Biochem. 2017 Jun;118(6):1432-1441. doi: 10.1002/jcb.25802. Epub 2017 Jan 10. J Cell Biochem. 2017. PMID: 27869314 Free PMC article.
-
Runx3 and Runx1 are required for CD8 T cell development during thymopoiesis.Proc Natl Acad Sci U S A. 2003 Jun 24;100(13):7731-6. doi: 10.1073/pnas.1232420100. Epub 2003 Jun 9. Proc Natl Acad Sci U S A. 2003. PMID: 12796513 Free PMC article.
-
Runx3 knockouts and stomach cancer.EMBO Rep. 2003 Jun;4(6):560-4. doi: 10.1038/sj.embor.embor868. EMBO Rep. 2003. PMID: 12776174 Free PMC article. Review.
References
-
- Ahn, M.-Y., S.-C. Bae, M. Maruyama, and Y. Ito. 1996. Comparison of the human genomic structure of the runt domain-encoding PEBP2/CBFα family. Gene 168:279-280. - PubMed
-
- Bae, S. C., E. Takahashi, Y. W. Zhang, E. Ogawa, K. Shigesada, Y. Namba, M. Satake, and Y. Ito. 1995. Cloning, mapping and expression of PEBP2 alpha C, a third gene encoding the mammalian Runt domain. Gene 159:245-248. - PubMed
-
- Banerjee, C., A. Javed, J. Y. Choi, J. Green, V. Rosen, A. J. van Wijnen, J. L. Stein, J. B. Lian, and G. S. Stein. 2001. Differential regulation of the two principal Runx2/Cbfa1 N-terminal isoforms in response to bone morphogenetic protein 2 during development of the osteoblast phenotype. Endocrinology 142:4026-4039. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

