Downstream sequence elements with different affinities for the hnRNP H/H' protein influence the processing efficiency of mammalian polyadenylation signals
- PMID: 11937639
- PMCID: PMC113221
- DOI: 10.1093/nar/30.8.1842
Downstream sequence elements with different affinities for the hnRNP H/H' protein influence the processing efficiency of mammalian polyadenylation signals
Abstract
Auxiliary factors likely play an important role in determining the polyadenylation efficiency of mammalian pre-mRNAs. We previously identified an auxiliary factor, hnRNP H/H', which stimulates 3'-end processing through an interaction with sequences downstream of the core elements of the SV40 late polyadenylation signal. Using in vitro reconstitution assays we have demonstrated that hnRNP H/H' can stimulate processing of two additional model polyadenylation signals by binding at similar relative downstream locations but with significantly different affinities. A short tract of G residues was determined to be a common property of all three hnRNP H/H' binding sites. A survey of mammalian polyadenylation signals identified potential G-rich hnRNP H/H' binding sites at similar downstream locations in approximately 34% of these signals. All of the novel G-rich elements tested were found to bind hnRNP H/H' protein and the processing of selected signals identified in the survey was stimulated by the protein both in vivo and in vitro. Downstream G-rich tracts, therefore, are a common auxiliary element in mammalian polyadenylation signals. Sequences capable of binding hnRNP H protein with varying affinities may play a role in determining the processing efficiency of a significant number of mammalian polyadenylation signals.
Figures




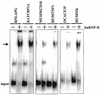


References
-
- Wahle E. and Ruegsegger,U. (1999) 3′-End processing of pre-mRNA in eukaryotes. FEMSMicrobiol. Rev., 23, 277–295. - PubMed

