Presence of a group II intron in a multiresistant Serratia marcescens strain that harbors three integrons and a novel gene fusion
- PMID: 11959575
- PMCID: PMC127176
- DOI: 10.1128/AAC.46.5.1402-1409.2002
Presence of a group II intron in a multiresistant Serratia marcescens strain that harbors three integrons and a novel gene fusion
Abstract
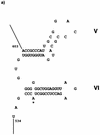
We analyzed the role of integrons in the dissemination of antibiotic resistance in a recent multiresistant clinical isolate, Serratia marcescens SCH88050909 (SCH909). This isolate harbors three integrons, all on a 60-kb conjugative plasmid. By PCR, hybridization, and sequencing analyses, we found that integron 1 has the dfrA1 and ant(3")-Ia cassettes. The first cassette in integron 2 contains the ant(2")-Ia gene, separated from its attC site (59-base element) by a 1,971-bp insert containing a group II intron; this intron codes for a putative maturase-reverse transcriptase on the complementary strand and is the first such intron to be found associated with an integron. The attC site is followed by a novel aminoglycoside resistance gene, ant(3")-Ii-aac(6')-IId, which has been characterized for its bifunctional ANT(3")-I and AAC(6')-II activities. DNA sequence analysis of this fused cassette suggests that insertion and excision due to the integrase activity could have an important role in the evolution of aminoglycoside resistance genes. This gene is followed by an unknown open reading frame with a typical attC site and a partial cassette composed of the beginning of the bla(OXA-10) cassette interrupted by IS1. The sequence downstream of IS1 revealed that the bla(OXA-10) cassette is incomplete and that the 3' conserved segment of this integron is absent. Integron 3 is in a Tn1696-like transposon with the aac(3)-Ia cassette followed by three unknown cassettes and ant(3")-Ia. The presence of the group II intron and the relationship of group II introns in eubacteria with mobile elements suggest a possible role of this element in events such as cassette formation and/or plasmid evolution.
Figures




Similar articles
-
The S.ma.I2 class C group II intron inserts at integron attC sites.Microbiology (Reading). 2008 May;154(Pt 5):1341-1353. doi: 10.1099/mic.0.2007/016360-0. Microbiology (Reading). 2008. PMID: 18451043
-
Potential role of group IIC-attC introns in integron cassette formation.J Bacteriol. 2009 Oct;191(19):6040-51. doi: 10.1128/JB.00674-09. Epub 2009 Jul 24. J Bacteriol. 2009. PMID: 19633079 Free PMC article.
-
Diversity and strength of internal outward-oriented promoters in group IIC-attC introns.Nucleic Acids Res. 2010 Dec;38(22):8196-207. doi: 10.1093/nar/gkq709. Epub 2010 Aug 16. Nucleic Acids Res. 2010. PMID: 20716518 Free PMC article.
-
Mobile gene cassettes and integrons: capture and spread of genes by site-specific recombination.Mol Microbiol. 1995 Feb;15(4):593-600. doi: 10.1111/j.1365-2958.1995.tb02368.x. Mol Microbiol. 1995. PMID: 7783631 Review.
-
Integrons: natural tools for bacterial genome evolution.Curr Opin Microbiol. 2001 Oct;4(5):565-9. doi: 10.1016/s1369-5274(00)00252-6. Curr Opin Microbiol. 2001. PMID: 11587934 Review.
Cited by
-
A structural analysis of the group II intron active site and implications for the spliceosome.RNA. 2010 Jan;16(1):1-9. doi: 10.1261/rna.1791310. Epub 2009 Nov 30. RNA. 2010. PMID: 19948765 Free PMC article.
-
Domain dissection and characterization of the aminoglycoside resistance enzyme ANT(3″)-Ii/AAC(6')-IId from Serratia marcescens.Biochimie. 2013 Jun;95(6):1319-25. doi: 10.1016/j.biochi.2013.02.011. Epub 2013 Feb 26. Biochimie. 2013. PMID: 23485681 Free PMC article.
-
Cosubstrate tolerance of the aminoglycoside resistance enzyme Eis from Mycobacterium tuberculosis.Antimicrob Agents Chemother. 2012 Nov;56(11):5831-8. doi: 10.1128/AAC.00932-12. Epub 2012 Sep 4. Antimicrob Agents Chemother. 2012. PMID: 22948873 Free PMC article.
-
Comprehensive review of chemical strategies for the preparation of new aminoglycosides and their biological activities.Chem Soc Rev. 2018 Feb 19;47(4):1189-1249. doi: 10.1039/c7cs00407a. Chem Soc Rev. 2018. PMID: 29296992 Free PMC article. Review.
-
Susceptibility of vertilmicin to modifications by three types of recombinant aminoglycoside-modifying enzymes.Antimicrob Agents Chemother. 2011 Aug;55(8):3950-3. doi: 10.1128/AAC.00300-11. Epub 2011 Jun 6. Antimicrob Agents Chemother. 2011. PMID: 21646489 Free PMC article.
References
-
- Cameron, F. H., D. J. Groot Obbink, V. P. Ackerman, and R. M. Hall. 1986. Nucleotide sequence of the AAD(2") aminoglycoside adenylyltransferase determinant aadB. Evolutionary relationship of this region with those surrounding aadA in R538-1 and dhfrII in R388. Nucleic Acids Res. 14:8625-8635. - PMC - PubMed
-
- Coffey, T. J., M. C. Enright, M. Daniels, J. K. Morona, R. Morona, W. Hryniewicz, J. C. Paton, and B. G. Spratt. 1998. Recombinational exchanges at the capsular polysaccharide biosynthetic locus lead to frequent serotype changes among natural isolates of Streptococcus pneumoniae. Mol. Microbiol. 27:73-83. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Medical

