The secE gene of Helicobacter pylori
- PMID: 11976315
- PMCID: PMC135042
- DOI: 10.1128/JB.184.10.2837-2840.2002
The secE gene of Helicobacter pylori
Abstract
Despite extensive annotation by two independent teams, the Helicobacter pylori genome appeared to lack a complete secretion machinery. The use of clinical isolates to substantiate in silico annotation is used here to identify the missing secE component of the major secretion machinery of Helicobacter pylori.
Figures


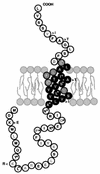
References
-
- Achtman, M., T. Azuma, D. E. Berg, Y. Ito, G. Morelli, Z. J. Pan, S. Suerbaum, S. A. Thompson, A. van der Ende, and L. J. van Doorn. 1999. Recombination and clonal groupings within Helicobacter pylori from different geographical regions. Mol. Microbiol. 32:459-470. - PubMed
-
- Bocs, S., A. Danchin, and C. Médigue. 2002. Re-annotation of genome microbial coding sequences: finding new genes and incorrectly annotated genes. BMC Bioinformatics 3:5. [Online.] http://www.biomedcentral.com/1471-2105/3/5. - PMC - PubMed
-
- Driessen, A. J., E. H. Manting, and C. van der Does. 2001. The structural basis of protein targeting and translocation in bacteria. Nat. Struct. Biol. 8:492-498. - PubMed
-
- Hartmann, E., T. Sommer, S. Prehn, D. Gorlich, S. Jentsch, and T. A. Rapoport. 1994. Evolutionary conservation of components of the protein translocation complex. Nature 367:654-657. - PubMed
-
- McIninch, J. D., W. S. Hayes, and M. Borodovsky. 1996. Applications of GeneMark in multispecies environments. Proc. Int. Conf. Intell. Syst. Mol. Biol. 4:165-175. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

