Evolution of bacteriophage in continuous culture: a model system to test antiviral gene therapies for the emergence of phage escape mutants
- PMID: 11992006
- PMCID: PMC137063
- DOI: 10.1128/jvi.76.11.5784-5792.2002
Evolution of bacteriophage in continuous culture: a model system to test antiviral gene therapies for the emergence of phage escape mutants
Abstract
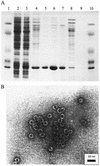
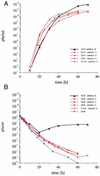
The emergence of viral escape mutants is usually a highly undesirable phenomenon. This phenomenon is frequently observed in antiviral drug applications for the treatment of viral infections and can undermine long-term therapeutic success. Here, we propose a strategy for evaluating a given antiviral approach in terms of its potential to provoke the appearance of resistant virus mutants. By use of Q beta RNA phage as a model system, the effect of an antiviral gene therapy, i.e., a virus-specific repressor protein expressed by a recombinant Escherichia coli host, was studied over the course of more than 100 generations. In 13 experiments carried out in parallel, 12 phage populations became resistant and 1 became extinct. Sequence analysis revealed that only two distinct phage mutants emerged in the 12 surviving phage populations. For both escape mutants, sequence variations located in the repressor binding site of the viral genomic RNA, which decrease affinity for the repressor protein, conferred resistance to translational repression. The results clearly suggest the feasibility of the proposed strategy for the evaluation of antiviral approaches in terms of their potential to allow resistant mutants to appear. In addition, the strategy proved to be a valuable tool for observing virus-specific molecular targets under the impact of antiviral drugs.
Figures
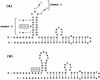
References
-
- Adams, M. H. 1959. Bacteriophages. Wiley Interscience, New York, N.Y.
-
- Arora, R., C. Priano, A. B. Jacobson, and D. R. Mills. 1996. cis-acting elements within an RNA coliphage genome: fold as you please, but fold you must. J. Mol. Biol. 258:433-446. - PubMed
-
- Ball, L. A., and P. Kaesberg. 1973. A polarity gradient in the expression of the replicase gene of RNA bacteriophage Qβ. J. Mol. Biol. 74:547-562. - PubMed
-
- Bauer, G., J. S. McCaskill, H. Otten, and A. Schwienhorst. 1989. Evolution im Laboratorium. Nachrbl. Chem. Tech. Lab. 37:484-488.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

