Identification of a novel cis-regulatory element involved in the heat shock response in Caenorhabditis elegans using microarray gene expression and computational methods
- PMID: 11997337
- PMCID: PMC186591
- DOI: 10.1101/gr.228902
Identification of a novel cis-regulatory element involved in the heat shock response in Caenorhabditis elegans using microarray gene expression and computational methods
Erratum in
- Genome Res 2002 Aug;12(8):1301
Abstract
We report here the identification of a previously unknown transcription regulatory element for heat shock (HS) genes in Caenorhabditis elegans. We monitored the expression pattern of 11,917 genes from C. elegans to determine the genes that were up-regulated on HS. Twenty eight genes were observed to be consistently up-regulated in several different repetitions of the experiments. We analyzed the upstream regions of these genes using computational DNA pattern recognition methods. Two potential cis-regulatory motifs were identified in this way. One of these motifs (TTCTAGAA) was the DNA binding motif for the heat shock factor (HSF), whereas the other (GGGTGTC) was previously unreported in the literature. We determined the significance of these motifs for the HS genes using different statistical tests and parameters. Comparative sequence analysis of orthologous HS genes from C. elegans and Caenorhabditis briggsae indicated that the identified DNA regulatory motifs are conserved across related species. The role of the identified DNA sites in regulation of HS genes was tested by in vitro mutagenesis of a green fluorescent protein (GFP) reporter transgene driven by the C. elegans hsp-16-2 promoter. DNA sites corresponding to both motifs are shown to play a significant role in up-regulation of the hsp-16-2 gene on HS. This is one of the rare instances in which a novel regulatory element, identified using computational methods, is shown to be biologically active. The contributions of individual sites toward induction of transcription on HS are nonadditive, which indicates interaction and cross-talk between the sites, possibly through the transcription factors (TFs) binding to these sites.
Figures

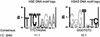



References
-
- Arnone MI, Davidson EH. The hardwiring of development: Organization and function of genomic regulatory systems. Development. 1997;124:1851–1864. - PubMed
-
- Berg OG, von Hippel PH. Selection of DNA binding sites by regulatory proteins: Statistical-mechanical theory and application to operators and promoters. J Mol Biol. 1987;193:723–750. - PubMed
-
- Bienz M, Pelham HRB. Mechanisms of heat shock gene activation in higher eukaryotes. Adv Genet. 1987;24:31–72. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
